
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 17,002,410 – 17,002,524 |
| Length | 114 |
| Max. P | 0.715040 |

| Location | 17,002,410 – 17,002,524 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -21.72 |
| Consensus MFE | -16.60 |
| Energy contribution | -16.85 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.715040 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 17002410 114 + 22407834 CUUUGGCAAUUAAUUGUUUAAUUUCGUUAUUUAAAAUUAAUGAUAUGUUUUUCAUAUGUCAUUGCACAUUUGUGCACUUAUUUUAUAUAAAUACAUGCCCGGUAUAUAUCCGUU ....(((((((((....)))))....((((.((((((.((((((((((.....))))))))))((((....))))....)))))).)))).....))))(((.......))).. ( -22.90) >DroSim_CAF1 111517 114 + 1 CUUUGGCAAUUAAUUGUUUAAUUUGGUUAUUUAAAAUUAAUGAUAUGUUUUUCAUAUGUCAUUGCACAUUUGUGCACUUAUUUUAUAUAAAUACAUGCCCGGUAUAUAUCCGUU ....(((((((((....))))).((.((((.((((((.((((((((((.....))))))))))((((....))))....)))))).))))...))))))(((.......))).. ( -23.60) >DroEre_CAF1 112131 114 + 1 CUUUGGCAAUUAAUUGUUUAAUUUGGUUAUUUAAAAUUAAUGAUAUGUUUUUCAUAUGUCAUUGCACAUUCGUGCACUUAUUUUAUAUAAAUACAUGUCCGGUAUAUAUCCGUU ....(((((((((....))))).((.((((.((((((.((((((((((.....))))))))))((((....))))....)))))).))))...))))))(((.......))).. ( -20.60) >DroAna_CAF1 101182 112 + 1 CUUUGGCAAUUAAUUGUUUAAUUUUGUUAUUUAAAAUUAAUGAUAUGUUUUUCAUAUGUCAUUGCACAUCUGUACGUGUAUUUUAUAUAAAUUCGCAUCAGGUACA--UGCGUU .....((((........(((((((((.....)))))))))((((((((.....))))))))))))..(((((...(((.((((.....)))).)))..)))))...--...... ( -19.80) >consensus CUUUGGCAAUUAAUUGUUUAAUUUGGUUAUUUAAAAUUAAUGAUAUGUUUUUCAUAUGUCAUUGCACAUUUGUGCACUUAUUUUAUAUAAAUACAUGCCCGGUAUAUAUCCGUU ....(((((((((....))))))...((((.((((((.((((((((((.....))))))))))((((....))))....)))))).))))......)))............... (-16.60 = -16.85 + 0.25)


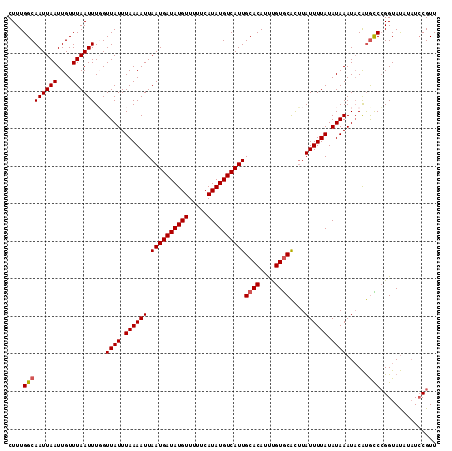
| Location | 17,002,410 – 17,002,524 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 92.11 |
| Mean single sequence MFE | -18.05 |
| Consensus MFE | -13.00 |
| Energy contribution | -13.12 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.541457 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
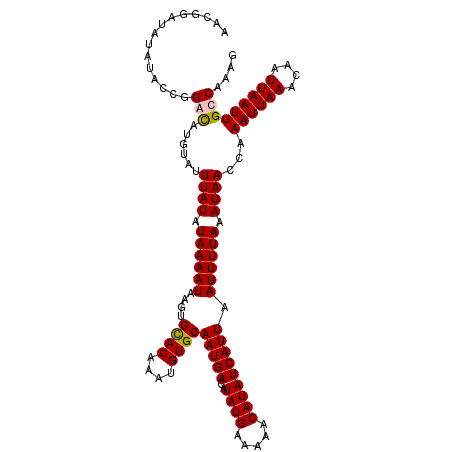
>2L_DroMel_CAF1 17002410 114 - 22407834 AACGGAUAUAUACCGGGCAUGUAUUUAUAUAAAAUAAGUGCACAAAUGUGCAAUGACAUAUGAAAAACAUAUCAUUAAUUUUAAAUAACGAAAUUAAACAAUUAAUUGCCAAAG ..(((.......)))(((((((.(((((((........(((((....))))).....)))))))..))))....((((((((.......))))))))..........))).... ( -20.62) >DroSim_CAF1 111517 114 - 1 AACGGAUAUAUACCGGGCAUGUAUUUAUAUAAAAUAAGUGCACAAAUGUGCAAUGACAUAUGAAAAACAUAUCAUUAAUUUUAAAUAACCAAAUUAAACAAUUAAUUGCCAAAG ..(((.......)))(((......((((.((((((....((((....))))(((((..((((.....))))))))).)))))).))))...((((((....))))))))).... ( -20.10) >DroEre_CAF1 112131 114 - 1 AACGGAUAUAUACCGGACAUGUAUUUAUAUAAAAUAAGUGCACGAAUGUGCAAUGACAUAUGAAAAACAUAUCAUUAAUUUUAAAUAACCAAAUUAAACAAUUAAUUGCCAAAG ..(((.......)))..(.(((((((((.....))))))))).)..((.(((((...(((((.....)))))..(((((((.........))))))).......)))))))... ( -15.90) >DroAna_CAF1 101182 112 - 1 AACGCA--UGUACCUGAUGCGAAUUUAUAUAAAAUACACGUACAGAUGUGCAAUGACAUAUGAAAAACAUAUCAUUAAUUUUAAAUAACAAAAUUAAACAAUUAAUUGCCAAAG ..((((--(.......)))))...((((.((((((....((((....))))(((((..((((.....))))))))).)))))).))))...((((((....))))))....... ( -15.60) >consensus AACGGAUAUAUACCGGACAUGUAUUUAUAUAAAAUAAGUGCACAAAUGUGCAAUGACAUAUGAAAAACAUAUCAUUAAUUUUAAAUAACCAAAUUAAACAAUUAAUUGCCAAAG ...............(((......((((.((((((....((((....))))(((((..((((.....))))))))).)))))).))))...((((((....))))))))).... (-13.00 = -13.12 + 0.12)

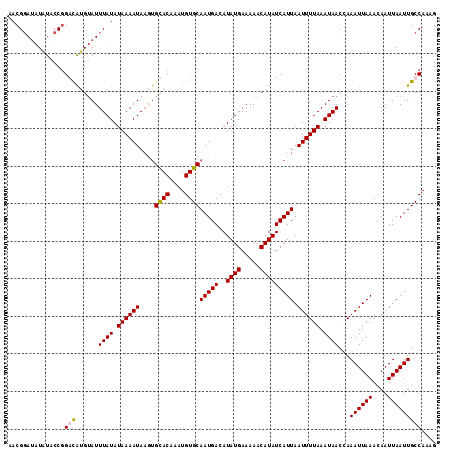
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:59:22 2006