
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,814,271 – 1,814,377 |
| Length | 106 |
| Max. P | 0.992572 |

| Location | 1,814,271 – 1,814,377 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.32 |
| Mean single sequence MFE | -23.65 |
| Consensus MFE | -11.46 |
| Energy contribution | -11.15 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.21 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.876834 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
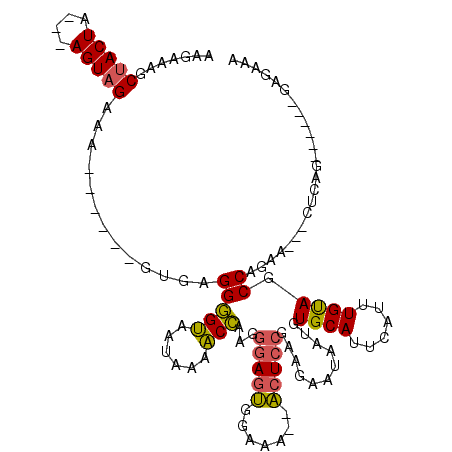
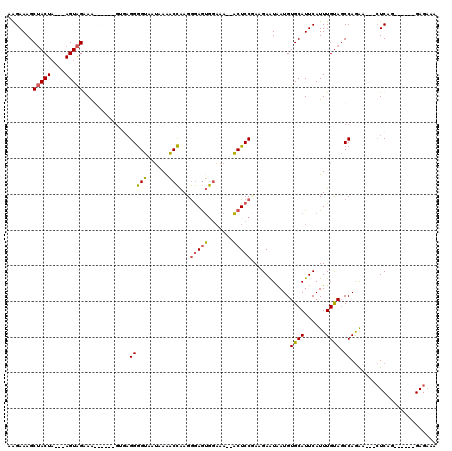
>2L_DroMel_CAF1 1814271 106 + 22407834 AAGAAAGCUACUA---AGUAGGAAGUGAUUGUGAGGGGUAAUAAAACCAAGGGAGUGGAAA--ACUCCGAAGAAUAAUGUGCAUUCAUUUGUAGCCAGGA---CUCAG------GAGAAA .......((((..---.)))).......((.((((.(((.(((((......(((((.....--)))))...((((.......)))).))))).)))....---)))).------)).... ( -21.70) >DroSim_CAF1 11929 106 + 1 AAGAAAGCUACUA---AGUAGGAAGUGUGCGUGAGGGGUAAUAAAACCAAGGGAGUGGAAA--ACUCCGAAGAAUAAUGUGCAUUCAUUUGUAGCCAAAA---GUCAG------GAGAAA ......(((((.(---(((.((..(((..(((....(((......)))...(((((.....--)))))........)))..)))))))))))))).....---.....------...... ( -24.10) >DroPer_CAF1 26332 108 + 1 AGGAAAGCAACUCAAAAGUGGAAG------GACAGGAGAGAGAAAUCUGAGAAAAGAAUAAGCUCUCCUAAGAGUGAUGUGCAUUCAUUUGUACCCAGGAGACCUCUG------GAGAAA ..(((.(((((((....((.....------.))(((((((.....(((......))).....)))))))..))))....))).)))........(((((.....))))------)..... ( -26.80) >DroEre_CAF1 22779 101 + 1 AAGAAAGCUACUA---AGUAGAAG----------GGGGUAAUAAGACCAAGGGAGUGGAAA--ACUCCCAAGAAUAAUGUGCAUUCAUUUGCAGCCAGAA---CUCAGAACCAGGAG-AA .......((((..---.))))..(----------(.(((......)))..((((((.....--))))))..........((((......)))).))....---(((........)))-.. ( -22.00) >consensus AAGAAAGCUACUA___AGUAGAAA______GUGAGGGGUAAUAAAACCAAGGGAGUGGAAA__ACUCCGAAGAAUAAUGUGCAUUCAUUUGUAGCCAGAA___CUCAG______GAGAAA .......(((((....))))).............(((((......)))...(((((.......)))))...........((((......)))).))........................ (-11.46 = -11.15 + -0.31)

| Location | 1,814,271 – 1,814,377 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.32 |
| Mean single sequence MFE | -19.56 |
| Consensus MFE | -11.66 |
| Energy contribution | -10.97 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.37 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.60 |
| SVM decision value | 2.34 |
| SVM RNA-class probability | 0.992572 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
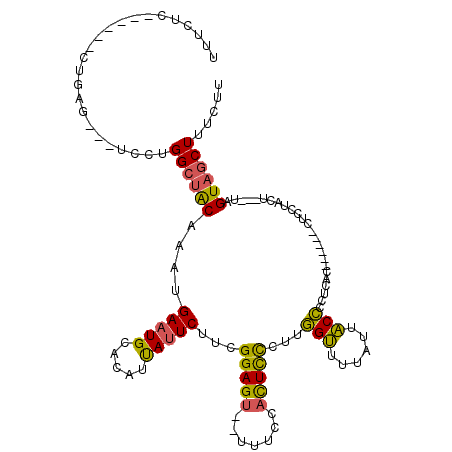
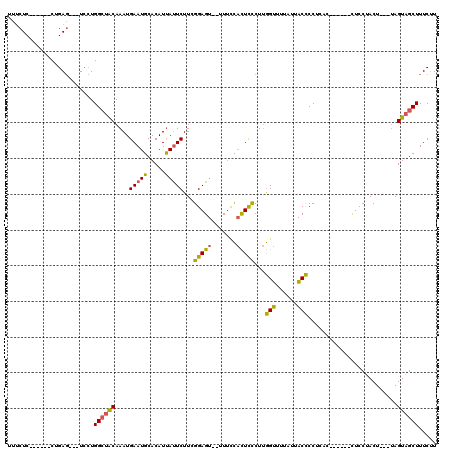
>2L_DroMel_CAF1 1814271 106 - 22407834 UUUCUC------CUGAG---UCCUGGCUACAAAUGAAUGCACAUUAUUCUUCGGAGU--UUUCCACUCCCUUGGUUUUAUUACCCCUCACAAUCACUUCCUACU---UAGUAGCUUUCUU ......------..(((---....((((((....(((((.....)))))...(((((--.....)))))...(((......)))....................---..))))))..))) ( -17.30) >DroSim_CAF1 11929 106 - 1 UUUCUC------CUGAC---UUUUGGCUACAAAUGAAUGCACAUUAUUCUUCGGAGU--UUUCCACUCCCUUGGUUUUAUUACCCCUCACGCACACUUCCUACU---UAGUAGCUUUCUU ......------.....---....((((((....(((((.....)))))...(((((--.....)))))...(((......)))....................---..))))))..... ( -16.30) >DroPer_CAF1 26332 108 - 1 UUUCUC------CAGAGGUCUCCUGGGUACAAAUGAAUGCACAUCACUCUUAGGAGAGCUUAUUCUUUUCUCAGAUUUCUCUCUCCUGUC------CUUCCACUUUUGAGUUGCUUUCCU ....((------(((.......))))).......(((.(((....((((.((((((((.....(((......))).....))))))))..------...........))))))).))).. ( -22.34) >DroEre_CAF1 22779 101 - 1 UU-CUCCUGGUUCUGAG---UUCUGGCUGCAAAUGAAUGCACAUUAUUCUUGGGAGU--UUUCCACUCCCUUGGUCUUAUUACCCC----------CUUCUACU---UAGUAGCUUUCUU ..-.....(((((((((---(...((.((((......))))......((..((((((--.....))))))..))...........)----------)....)))---))).))))..... ( -22.30) >consensus UUUCUC______CUGAG___UCCUGGCUACAAAUGAAUGCACAUUAUUCUUCGGAGU__UUUCCACUCCCUUGGUUUUAUUACCCCUCAC______CUCCUACU___UAGUAGCUUUCUU ........................((((((....(((((.....)))))...(((((.......)))))...(((......))).........................))))))..... (-11.66 = -10.97 + -0.69)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:45 2006