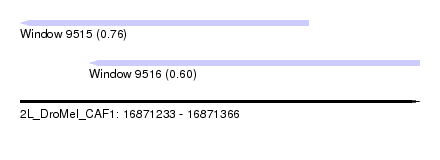

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,871,233 – 16,871,366 |
| Length | 133 |
| Max. P | 0.757444 |

| Location | 16,871,233 – 16,871,329 |
|---|---|
| Length | 96 |
| Sequences | 3 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 65.75 |
| Mean single sequence MFE | -27.29 |
| Consensus MFE | -11.02 |
| Energy contribution | -13.91 |
| Covariance contribution | 2.89 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.40 |
| SVM decision value | 0.49 |
| SVM RNA-class probability | 0.757444 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
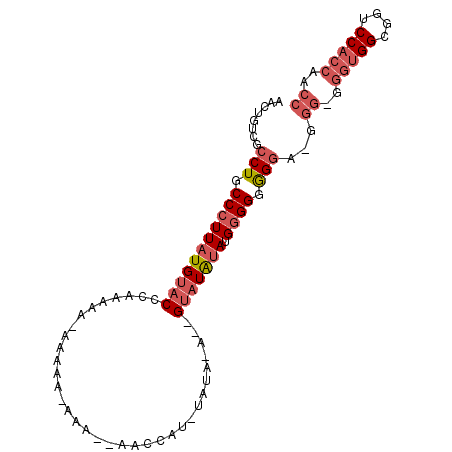
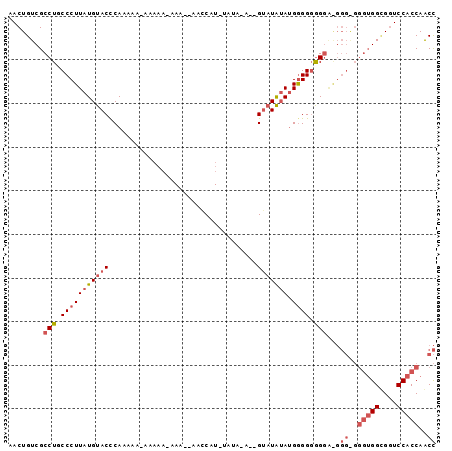
>2L_DroMel_CAF1 16871233 96 - 22407834 AACUGUCGCCUGCCCUUAUGUACCCAAAAAGAAAAAGAAA--AACCAUGUAUAUAUGGGUUGGAUGAGGGAGGAAGGGAGGGUGGCGGUCCACCAGCC .((((((((((.(((((.....(((...............--..(((((....))))).((....)))))...))))).))))))))))......... ( -29.30) >DroSec_CAF1 10559 88 - 1 AACUGUCGCCUGCCCUUAUGUACUCAAAAA--------AA--AACCAUAUAUAAACCGUAUAUAAGGGGGGGGGGGGGGGGGUGGUGCUCCACCAACC ..((.((.(((.(((((((((((.......--------..--...............))))))))))).))).)).))..(((((....))))).... ( -34.26) >DroEre_CAF1 42842 80 - 1 AACUGUCGCCUGCCCUUAUGUACCCAAAAAAAAAAAAAAAAUAAACGU---------GUAUAUAUGGGGUGGUU---------GGGGGUCCACCAACU .(((.(((.(..(((((((((((.(.....................).---------))))))).))))..).)---------)).)))......... ( -18.30) >consensus AACUGUCGCCUGCCCUUAUGUACCCAAAAA_AAAAA_AAA__AACCAU_UAUA_A__GUAUAUAUGGGGGGGGA_GGG_GGGUGGCGGUCCACCAACC ........(((.(((((((((((..................................))))))).)))).)))...((..(((((....)))))..)) (-11.02 = -13.91 + 2.89)

| Location | 16,871,256 – 16,871,366 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 80.34 |
| Mean single sequence MFE | -23.66 |
| Consensus MFE | -13.42 |
| Energy contribution | -14.86 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.57 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.598164 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
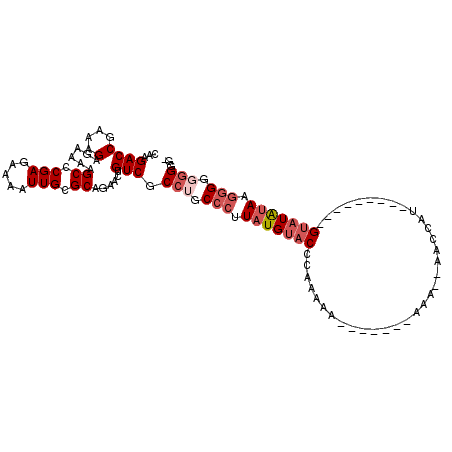
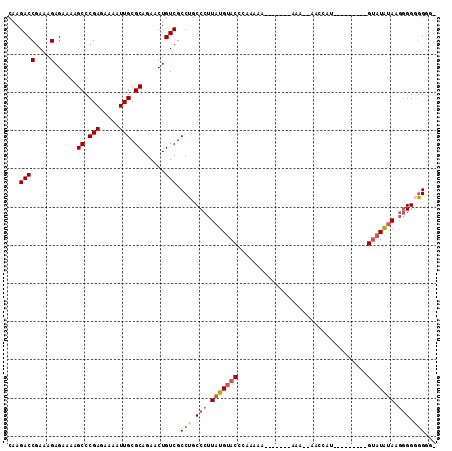
>2L_DroMel_CAF1 16871256 110 - 22407834 CAAGACCGAAAGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACCCAAAAAGAAAAAGAAA--AACCAUGUAUAUAUGGGUUGGAUGAGGGAGGAA ...((((....)......((.(((.....))).))......))).(((.(((((((.((((((..............--.............)))).)).)))))))))).. ( -25.33) >DroSec_CAF1 10582 102 - 1 CAAGACCGAAAGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACUCAAAAA--------AA--AACCAUAUAUAAACCGUAUAUAAGGGGGGGGGG ......(....).......(((.........(((((....)).)))((.(((((((((((.......--------..--...............))))))))))).))))). ( -26.66) >DroSim_CAF1 13113 93 - 1 CAAGACCGAAAGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACUCAAAAA-------AAA--AACCAU---------GUAUAUAAGGGGGGGAG- ...((((....)......((.(((.....))).))......))).(((.(((((((((((.......-------...--......---------))))))))))).)))..- ( -27.29) >DroEre_CAF1 42857 102 - 1 CAAGACCGAAGGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACCCAAAAAAAAAAAAAAAAUAAACGU---------GUAUAUAUGGGGUGGUU- ...((((..(((.((...((.(((.....))).)).......)).)))((((((((((((.(.....................).---------))))))).)))))))))- ( -19.50) >DroYak_CAF1 5978 94 - 1 CAAGACCGAAAGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACCCAAAA--------AAAAUAAACGU---------GUAUAUACUGGGUGGUU- ...((((....)......((.(((.....))).))......)))(((.((((.(((((((.(....--------.........).---------)))))))..))))))).- ( -19.52) >consensus CAAGACCGAAAGAGAAAAGCCCGAGAAAAUUGCGCAGAACUGUCGCCUGCCCUUAUGUACCCAAAAA_______AAA__AACCAU_________GUAUAUAAGGGGGGGGG_ ...((((....)......((.(((.....))).))......))).(((.(((.(((((((..................................))))))).))).)))... (-13.42 = -14.86 + 1.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:58:17 2006