
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,801,028 – 16,801,124 |
| Length | 96 |
| Max. P | 0.863823 |

| Location | 16,801,028 – 16,801,124 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 78.46 |
| Mean single sequence MFE | -18.10 |
| Consensus MFE | -13.78 |
| Energy contribution | -13.25 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.863823 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16801028 96 - 22407834 CU------CGUGCCCGUGUAGCUUAAUGACAGAAAUUAGGCAACCUUUCACUGGCACAUAAUUUCACCCACACACA-CACACUCGCACUUGCAA--UGACUUAUA ..------.(((((.(((..(((((((.......))))))).......))).)))))...................-.......((....))..--......... ( -19.20) >DroPse_CAF1 3548 92 - 1 CU------CGUGCUCGUGUCGUUUAAUGACAGAAAUUAGGCAACCUUUCACUGGCGCAUAAUUUCACCCACACGCA-CAC------ACUCGCACUCGUCCCCAUA ..------.((((...((((((...))))))(((((((.((..((.......)).)).)))))))........)))-)..------................... ( -19.20) >DroEre_CAF1 3011 89 - 1 CU------CGUGCCCGUGUAGCUUAAUGACUGAAAUUAGGCAACCUUUCACUGGCACAUAGUUUCACCCACACACA-UAG------ACU-GCAG--CGACUUAUA ..------.(((((.(((..(((((((.......))))))).......))).))))).((((((............-.))------)))-)...--......... ( -18.32) >DroYak_CAF1 3036 90 - 1 CU------CGUGCCCGUGUAGCUUAAUGACAGAAAUUAGGCAACCUUUCACUGGCACAUAAUUUCACCCACACACA-UAC------ACUCGCAG--UGACAUAUA ..------.(((((.(((..(((((((.......))))))).......))).)))))...................-..(------((.....)--))....... ( -19.60) >DroAna_CAF1 3118 99 - 1 UUCCUAUUCGUGAUGGAGUAGUUUAAUGACAAAAAUUAGGCAACCUUUCACUGCCAUACACUCUCACUUGAAGCCAAAAC------AAUGACAGCCUGACAUAUA .........((((.((.((((((....)))........((((.........)))).))).)).)))).............------................... ( -13.10) >DroPer_CAF1 3594 92 - 1 CU------CGUGCUCGUGUCGUUUAAUGACAGAAAUUAGGCAACCUUUCACUGGCGCAUAAUUUCACCCACACGCA-CAC------ACUCGCACUCGUCCCCAUA ..------.((((...((((((...))))))(((((((.((..((.......)).)).)))))))........)))-)..------................... ( -19.20) >consensus CU______CGUGCCCGUGUAGCUUAAUGACAGAAAUUAGGCAACCUUUCACUGGCACAUAAUUUCACCCACACACA_CAC______ACUCGCAG__UGACAUAUA .........(((((.(((..(((((((.......))))))).......))).)))))................................................ (-13.78 = -13.25 + -0.53)

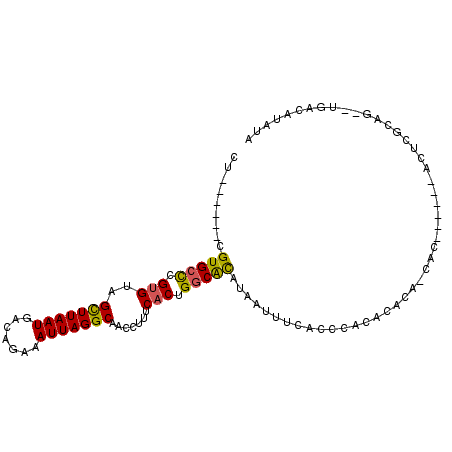
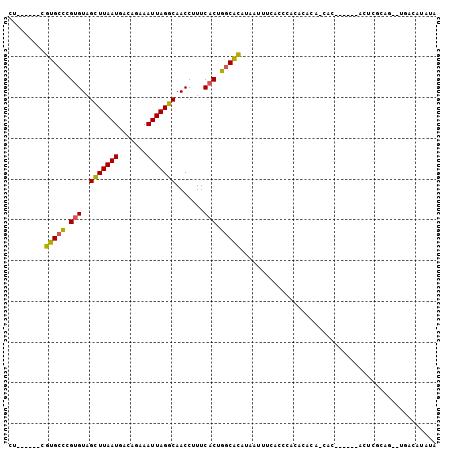
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:47 2006