
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,770,876 – 16,770,972 |
| Length | 96 |
| Max. P | 0.964192 |

| Location | 16,770,876 – 16,770,972 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 81.79 |
| Mean single sequence MFE | -33.52 |
| Consensus MFE | -23.43 |
| Energy contribution | -22.85 |
| Covariance contribution | -0.58 |
| Combinations/Pair | 1.28 |
| Mean z-score | -4.12 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.56 |
| SVM RNA-class probability | 0.964192 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
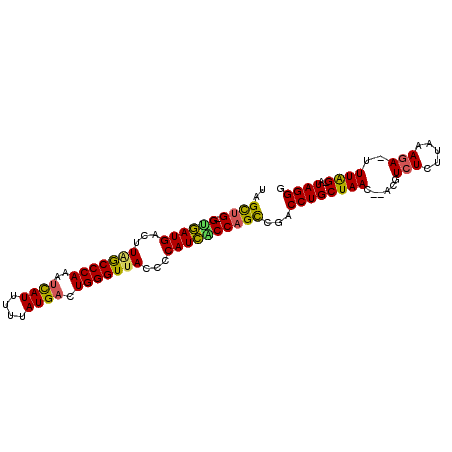
>2L_DroMel_CAF1 16770876 96 + 22407834 UAGCUG-GUGAUGACUUAGCCCAAAUCAUUUUUAUGACUGGGUUACCCCAUCACCAGUCGACCUGCUAACACACGUCUCUUAAAGA-UUUAG-AUAGGG ..((((-((((((...(((((((..((((....)))).)))))))...))))))))))...((((((((.....((((.....)))-)))))-.)))). ( -33.90) >DroSec_CAF1 498 95 + 1 UAGCUG-GUGAUGACUUGGCCCAAAUCAUUUUUAUGACUGGGUUACGCCAUCACCAGUCGACCUGCUAAC-CACGUCUCUUAAAGA-UUUAG-AUAGGG ..((((-((((((.(.(((((((..((((....)))).))))))).).))))))))))...((((((((.-...((((.....)))-)))))-.)))). ( -35.10) >DroSim_CAF1 294 96 + 1 UAGUUG-GUGAUGACUUGGCCCAAAUCAUUUUAAUGACUGGGUUACGCCAUCACCAGCCGACCUGCUAACACAUGUCUCUUAAAGA-UUUAG-AUAGGG ..((((-((((((.(.(((((((..((((....)))).))))))).).))))))))))...((((((((.....((((.....)))-)))))-.)))). ( -33.80) >DroEre_CAF1 498 95 + 1 CAGCUGUGGGAUGACUUAGCCCAACUCAUUUUUAUGACUGGGUUACCGCAUCCCCAGCCGACCUGCUAAC--ACGUCUCUUAAAGA-UUUGG-AUAGGG ..((((.((((((...(((((((..((((....)))).)))))))...))))))))))...((((((((.--..((((.....)))-)))))-.)))). ( -36.10) >DroYak_CAF1 516 94 + 1 UAGCUG-GUGAUGACUUAGCCCAAAAUAUUUUUAUGACUGGGUUAUCUCAUCCCCAGCCGACCUGCUAAC--ACGUCUCUUAAAGA-UUUAG-AUAGGG ..((((-(.(((((..(((((((..(((....)))...)))))))..))))).)))))...((((((((.--..((((.....)))-)))))-.)))). ( -33.90) >DroAna_CAF1 506 83 + 1 UAGCUGGGAAAUGGCUUAACCCA--------------UUGGGUGAGUUCAUUUCCCGCUGACCUGCUAAC--ACGUCUCUCAAACAGUUUAGUUUAGGG ((((.(((((((((((((.(((.--------------..))))))).))))))))))))).((((((((.--((............)))))))..))). ( -28.30) >consensus UAGCUG_GUGAUGACUUAGCCCAAAUCAUUUUUAUGACUGGGUUACCCCAUCACCAGCCGACCUGCUAAC__ACGUCUCUUAAAGA_UUUAG_AUAGGG ..((((.((((((...(((((((..((((....)))).)))))))...))))))))))...((((((((......(((.....)))..))))..)))). (-23.43 = -22.85 + -0.58)



| Location | 16,770,876 – 16,770,972 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 81.79 |
| Mean single sequence MFE | -31.67 |
| Consensus MFE | -19.52 |
| Energy contribution | -20.47 |
| Covariance contribution | 0.95 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.33 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929618 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16770876 96 - 22407834 CCCUAU-CUAAA-UCUUUAAGAGACGUGUGUUAGCAGGUCGACUGGUGAUGGGGUAACCCAGUCAUAAAAAUGAUUUGGGCUAAGUCAUCAC-CAGCUA ......-.....-.........(((.(((....))).)))(.((((((((((..((.(((((((((....)))).))))).))..)))))))-)))).. ( -34.10) >DroSec_CAF1 498 95 - 1 CCCUAU-CUAAA-UCUUUAAGAGACGUG-GUUAGCAGGUCGACUGGUGAUGGCGUAACCCAGUCAUAAAAAUGAUUUGGGCCAAGUCAUCAC-CAGCUA ......-.....-.........(((.((-.....)).)))(.(((((((((((....(((((((((....)))).)))))....))))))))-)))).. ( -34.10) >DroSim_CAF1 294 96 - 1 CCCUAU-CUAAA-UCUUUAAGAGACAUGUGUUAGCAGGUCGGCUGGUGAUGGCGUAACCCAGUCAUUAAAAUGAUUUGGGCCAAGUCAUCAC-CAACUA ......-.....-.........(((.(((....))).)))((.((((((((((....(((((((((....)))).)))))....))))))))-)).)). ( -32.00) >DroEre_CAF1 498 95 - 1 CCCUAU-CCAAA-UCUUUAAGAGACGU--GUUAGCAGGUCGGCUGGGGAUGCGGUAACCCAGUCAUAAAAAUGAGUUGGGCUAAGUCAUCCCACAGCUG ......-.....-.........(((.(--(....)).)))((((((((((((..((.(((((((((....)))).))))).)).).)))))).))))). ( -34.10) >DroYak_CAF1 516 94 - 1 CCCUAU-CUAAA-UCUUUAAGAGACGU--GUUAGCAGGUCGGCUGGGGAUGAGAUAACCCAGUCAUAAAAAUAUUUUGGGCUAAGUCAUCAC-CAGCUA ......-.....-.........(((.(--(....)).)))((((((.(((((..((.(((((.............))))).))..))))).)-))))). ( -29.02) >DroAna_CAF1 506 83 - 1 CCCUAAACUAAACUGUUUGAGAGACGU--GUUAGCAGGUCAGCGGGAAAUGAACUCACCCAA--------------UGGGUUAAGCCAUUUCCCAGCUA ...(((((......)))))...(((.(--(....)).)))(((((((((((.....((((..--------------.)))).....)))))))).))). ( -26.70) >consensus CCCUAU_CUAAA_UCUUUAAGAGACGU__GUUAGCAGGUCGGCUGGUGAUGGGGUAACCCAGUCAUAAAAAUGAUUUGGGCUAAGUCAUCAC_CAGCUA ......................(((.(........).)))((((((((((((.....(((((((((....)))).))))).....))))))).))))). (-19.52 = -20.47 + 0.95)


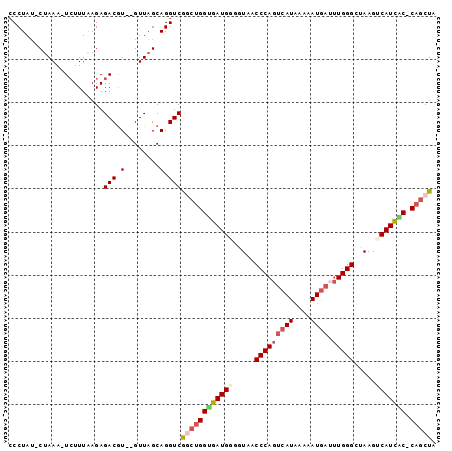
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:57:10 2006