
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,708,202 – 16,708,315 |
| Length | 113 |
| Max. P | 0.689237 |

| Location | 16,708,202 – 16,708,315 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.57 |
| Mean single sequence MFE | -29.33 |
| Consensus MFE | -19.12 |
| Energy contribution | -19.69 |
| Covariance contribution | 0.57 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.597882 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16708202 113 + 22407834 CUUUGAAUGAGCUGAAAUUAAAGUUGAAUUGAAAAGCAAGCAAAGUGCUUAGGAAUUCAAAUAAUU--GGGAAU-----UGCCUUAGGCAAUUAGUAAAUUCAUUUCAGUCAUUAAGGGC ((((((.(((.(((((((.....(((((((...(((((.......)))))...)))))))....((--(..(((-----((((...)))))))..)))....)))))))))))))))).. ( -28.00) >DroSec_CAF1 162788 113 + 1 CUCGGAAUGAGCUGAAAUUAAAGUUGAAUUGAAGGGCAAGCAAAGUGCUUAGGAAUUCAAAUAAUU--GGGAAU-----UGCCUUAGGCAAUUAGUAAAUUCAUUUCAGCCAUUAAGGGC (((..((((.((((((((.....(((((((.(((.((.......)).)))...)))))))....((--(..(((-----((((...)))))))..)))....))))))))))))..))). ( -31.20) >DroEre_CAF1 163957 120 + 1 CUCGGAAUUAGCUGCAAUUAAAGUUGGAUUCAAGGGCUGGCAUGGUGCUUUGGAAUUGAAAUUGUUUUUGGAAUGGAGAAGCCUCAGGCAAUUGGUAAAUUCAUUUCAGCCAUUAAGGCC ((.(((.((((((........)))))).))).))((((...(((((......(((((..((((((((..((..........))..))))))))....)))))......)))))...)))) ( -28.80) >consensus CUCGGAAUGAGCUGAAAUUAAAGUUGAAUUGAAGGGCAAGCAAAGUGCUUAGGAAUUCAAAUAAUU__GGGAAU_____UGCCUUAGGCAAUUAGUAAAUUCAUUUCAGCCAUUAAGGGC (((..((((.((((((((.....(((((((...((((.........))))...)))))))..........((((.....((((...))))........))))))))))))))))..))). (-19.12 = -19.69 + 0.57)


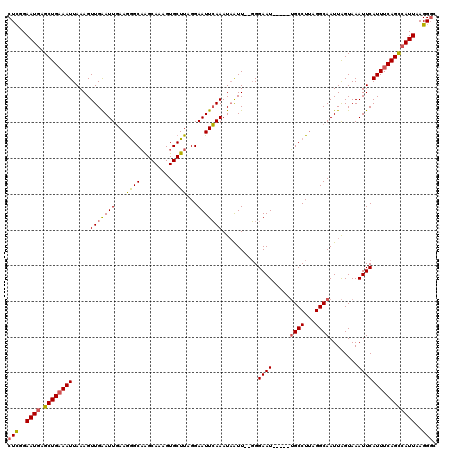
| Location | 16,708,202 – 16,708,315 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.57 |
| Mean single sequence MFE | -21.83 |
| Consensus MFE | -14.36 |
| Energy contribution | -15.92 |
| Covariance contribution | 1.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.689237 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16708202 113 - 22407834 GCCCUUAAUGACUGAAAUGAAUUUACUAAUUGCCUAAGGCA-----AUUCCC--AAUUAUUUGAAUUCCUAAGCACUUUGCUUGCUUUUCAAUUCAACUUUAAUUUCAGCUCAUUCAAAG ......((((((((((((.........(((((((...))))-----)))...--......(((((((...(((((.......)))))...))))))).....))))))).)))))..... ( -24.50) >DroSec_CAF1 162788 113 - 1 GCCCUUAAUGGCUGAAAUGAAUUUACUAAUUGCCUAAGGCA-----AUUCCC--AAUUAUUUGAAUUCCUAAGCACUUUGCUUGCCCUUCAAUUCAACUUUAAUUUCAGCUCAUUCCGAG ...(((((((((((((((.........(((((((...))))-----)))...--......(((((((..(((((.....)))))......))))))).....)))))))).))))..))) ( -24.50) >DroEre_CAF1 163957 120 - 1 GGCCUUAAUGGCUGAAAUGAAUUUACCAAUUGCCUGAGGCUUCUCCAUUCCAAAAACAAUUUCAAUUCCAAAGCACCAUGCCAGCCCUUGAAUCCAACUUUAAUUGCAGCUAAUUCCGAG ((..((((.(((((...((.......))...((.((..((((............................))))..)).))))))).))))..))......................... ( -16.49) >consensus GCCCUUAAUGGCUGAAAUGAAUUUACUAAUUGCCUAAGGCA_____AUUCCC__AAUUAUUUGAAUUCCUAAGCACUUUGCUUGCCCUUCAAUUCAACUUUAAUUUCAGCUCAUUCCGAG .........(((((((((((((........((((...)))).....))))..........(((((....(((((.....)))))...)))))..........)))))))))......... (-14.36 = -15.92 + 1.56)

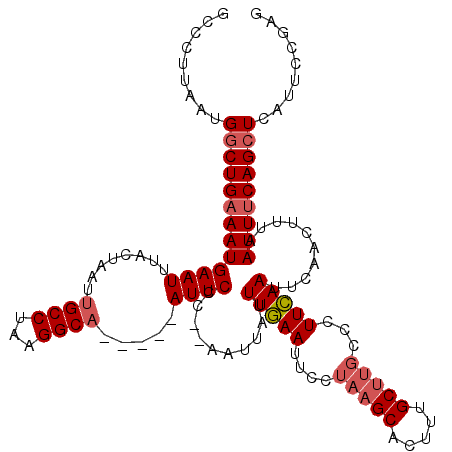
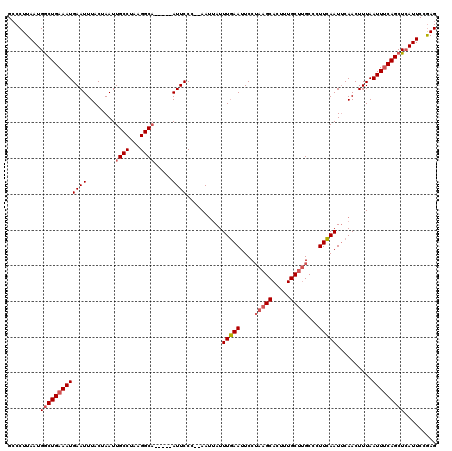
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:41 2006