
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,706,959 – 16,707,159 |
| Length | 200 |
| Max. P | 0.978509 |

| Location | 16,706,959 – 16,707,079 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.56 |
| Mean single sequence MFE | -37.53 |
| Consensus MFE | -35.37 |
| Energy contribution | -36.37 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.88 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16706959 120 + 22407834 GAACAGCUGGAAUGAGUUGCAAACAUCACAGACAAUGCCGUUUAAAUGGAAAAUUGUUAGACGUCACAGGCCAGCGAGAGCUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUC ((((((((((..((.(((((..........((((((.((((....))))...)))))).(((.....(((..(((....)))....))).....)))..)))))))....)))))))))) ( -41.50) >DroSec_CAF1 161606 120 + 1 GAACAGCUGGAAUGAGUUGCAAACAUCACAGACAAUGCCGUUUAAAUGGAAAAUUGUUAGACGUCACAGGCCAGCAAGAGCUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUC ((((((((((..((.(((((..........((((((.((((....))))...)))))).(((.....(((..(((....)))....))).....)))..)))))))....)))))))))) ( -38.80) >DroEre_CAF1 162766 120 + 1 GAACAGCUGGAAUGAGUUGCGAACAUCACAGACAAUGCCUUUUAAAUGGAAAAUUGUUAGACGUCACAGGCCAGCGAUAGCUACAACCUAAGCAGUCCGAAGACCAGUUGUCAGCUGUUC ((((((((((..((.(((.((.........((((((.((........))...)))))).(((.....(((..(((....)))....))).....)))))..)))))....)))))))))) ( -32.30) >consensus GAACAGCUGGAAUGAGUUGCAAACAUCACAGACAAUGCCGUUUAAAUGGAAAAUUGUUAGACGUCACAGGCCAGCGAGAGCUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUC ((((((((((..((.(((((..........((((((.((((....))))...)))))).(((.....(((..(((....)))....))).....)))..)))))))....)))))))))) (-35.37 = -36.37 + 1.00)

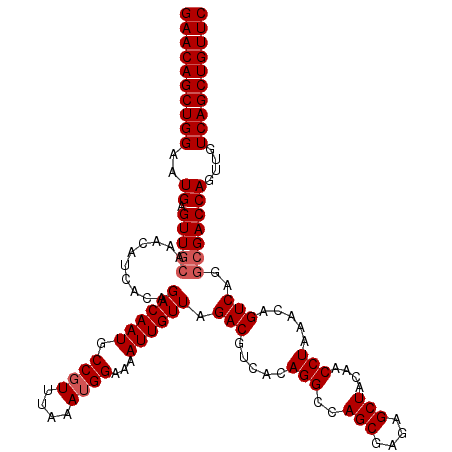
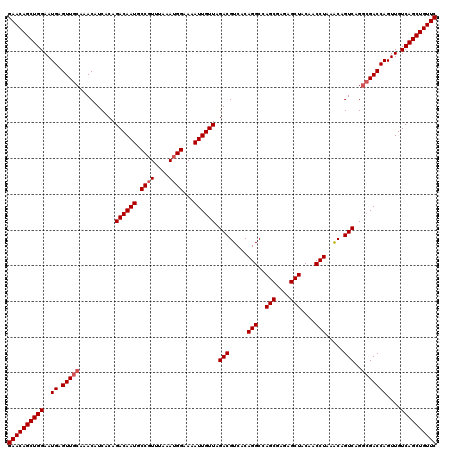
| Location | 16,706,959 – 16,707,079 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.56 |
| Mean single sequence MFE | -36.47 |
| Consensus MFE | -34.49 |
| Energy contribution | -34.27 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.02 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.95 |
| SVM decision value | 0.88 |
| SVM RNA-class probability | 0.872509 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16706959 120 - 22407834 GAACAGCUGACAACUGGUCGCCUGACUGUUUAGGUUGUAGCUCUCGCUGGCCUGUGACGUCUAACAAUUUUCCAUUUAAACGGCAUUGUCUGUGAUGUUUGCAACUCAUUCCAGCUGUUC (((((((((......((((((((((....))))))...(((....)))))))((..(((((..(((((...((........)).)))))....)))).)..))........))))))))) ( -35.20) >DroSec_CAF1 161606 120 - 1 GAACAGCUGACAACUGGUCGCCUGACUGUUUAGGUUGUAGCUCUUGCUGGCCUGUGACGUCUAACAAUUUUCCAUUUAAACGGCAUUGUCUGUGAUGUUUGCAACUCAUUCCAGCUGUUC (((((((((......((((((((((....))))))...(((....)))))))((..(((((..(((((...((........)).)))))....)))).)..))........))))))))) ( -35.80) >DroEre_CAF1 162766 120 - 1 GAACAGCUGACAACUGGUCUUCGGACUGCUUAGGUUGUAGCUAUCGCUGGCCUGUGACGUCUAACAAUUUUCCAUUUAAAAGGCAUUGUCUGUGAUGUUCGCAACUCAUUCCAGCUGUUC (((((((((((((((((((....)))).....)))))).((((....)))).(((((((((..(((((...((........)).)))))....)))).)))))........))))))))) ( -38.40) >consensus GAACAGCUGACAACUGGUCGCCUGACUGUUUAGGUUGUAGCUCUCGCUGGCCUGUGACGUCUAACAAUUUUCCAUUUAAACGGCAUUGUCUGUGAUGUUUGCAACUCAUUCCAGCUGUUC (((((((((((((((((((....)))).....))))))(((....)))((..(((((((((..(((((...((........)).)))))....)))).)))))..))....))))))))) (-34.49 = -34.27 + -0.22)


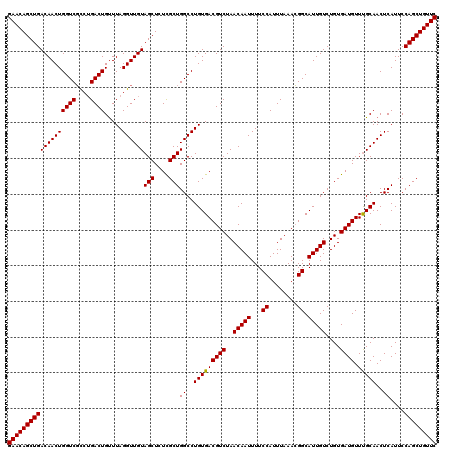
| Location | 16,707,039 – 16,707,159 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.43 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -30.81 |
| Energy contribution | -31.03 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.62 |
| Structure conservation index | 1.02 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870950 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16707039 120 + 22407834 CUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUCGGGGAAAACUUUACGCUGCACCGCACAAAAUCACCAAACAAUGGCCCACAAUUAGGUUGGCCCUUAAGUCAAGUUCCUUU ......((..((((((..(((((....))))).))))))..))...(((((.((((......))..................((((.((......)).)))).....)).)))))..... ( -28.80) >DroSec_CAF1 161686 119 + 1 CUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUCGGGGAAAACUUUACGCUGCACCGCACAAAAUC-CCAAACAAUGGCCAACAAUUAGGUUGGCCCUUAAGUCAAGUUCCUUU ......((..((((((..(((((....))))).))))))..))...(((((.((((......))........-.........(((((((......))))))).....)).)))))..... ( -32.90) >DroEre_CAF1 162846 117 + 1 CUACAACCUAAGCAGUCCGAAGACCAGUUGUCAGCUGUUCGGGGAAAACUUUACGCUGCACCGCACAAAAUC-CAACACAAUGGCCAACAAUUAGGUUGGCCCUUAAGUCAAGUUCCU-- ......((..((((((..((.((....)).)).))))))..))...(((((.((((......))........-.........(((((((......))))))).....)).)))))...-- ( -28.60) >consensus CUACAACCUAAACAGUCAGGCGACCAGUUGUCAGCUGUUCGGGGAAAACUUUACGCUGCACCGCACAAAAUC_CCAAACAAUGGCCAACAAUUAGGUUGGCCCUUAAGUCAAGUUCCUUU ......((..((((((..(((((....))))).))))))..))...(((((.((((......))..................(((((((......))))))).....)).)))))..... (-30.81 = -31.03 + 0.23)

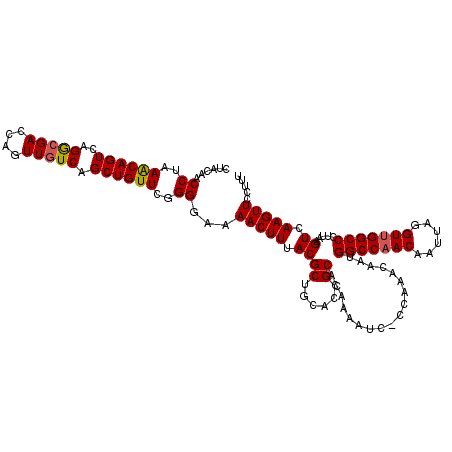
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:36 2006