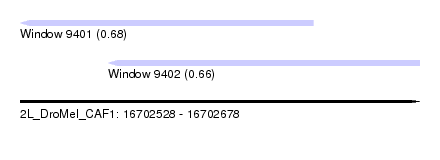

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,702,528 – 16,702,678 |
| Length | 150 |
| Max. P | 0.683936 |

| Location | 16,702,528 – 16,702,638 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 86.21 |
| Mean single sequence MFE | -30.05 |
| Consensus MFE | -22.39 |
| Energy contribution | -23.82 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683936 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16702528 110 - 22407834 AAAUAUGGGA---GGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUCCUUCCUCAAUUCUAUGUAAUCUUCGCUGCCAGGGGCAU------- ..((((((((---((((.((((((((((((((((((((....)).)))))...((.....)))))))))))).)))))))))....)))))).........((((...)))).------- ( -33.80) >DroSec_CAF1 157232 113 - 1 AAAUAUGGGAGGAGGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUUCUUCCUCAAUUCUAUGUAAUCUUUGCUGCCAGGUAAAU------- ..((((((((.((((((.((((((((((((((((((((....)).)))))...((.....)))))))))))).)))))))))..))))))))....((((((....)))))).------- ( -34.30) >DroSim_CAF1 157157 113 - 1 AAAUAUGGGAGGAGGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAUUGACAUGGAACACACGCAAUCUGUCUCCUUCCUCAAUUCUAUGUAAUCUUUGCUGCCAGGUACAU------- ..((((((((.((((((.(((((((((((((...((......((((...)))).....))..)))))))))).)))))))))..))))))))......((((....))))...------- ( -32.20) >DroEre_CAF1 158605 114 - 1 AAAUGUGGGC---GGGUUGGAAUAGAUUGUGCAUUGCAUAAUUACCAAUGGCACGGAACACACGCAAUCUUUCU---UCUUCAAUUCUAUGUAAUCUUUUCUACCAGGGACAUACUCGUU .......(((---(((((((((.(((((((((((((.........)))))....(.....).))))))))))).---........(((..(((........)))...))))).))))))) ( -19.90) >consensus AAAUAUGGGA___GGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUCCUUCCUCAAUUCUAUGUAAUCUUUGCUGCCAGGGACAU_______ ..(((((((..((((((.((((((((((((((((((((....)).)))))....(.....).)))))))))).)))))))))...)))))))......((((....)))).......... (-22.39 = -23.82 + 1.44)


| Location | 16,702,561 – 16,702,678 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.52 |
| Mean single sequence MFE | -34.25 |
| Consensus MFE | -26.12 |
| Energy contribution | -27.62 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.76 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.659718 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16702561 117 - 22407834 GAGAAUUGAAAUUAAUUUGCUGCCAGAAGGCGGUGGGUGGAAAUAUGGGA---GGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUCCUUCC ..............(((..(((((....)))))..)))........((((---((((....(((((((((((((((((....)).)))))...((.....)))))))))))))))))))) ( -39.40) >DroSec_CAF1 157265 120 - 1 GGGAAUUGAAAUUAAUUUGCUGCCAGAAGGCGGUGGGUGGAAAUAUGGGAGGAGGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUUCUUCC ..............(((..(((((....)))))..))).........(((((((((......((((((((((((((((....)).)))))...((.....)))))))))))))))))))) ( -36.60) >DroSim_CAF1 157190 120 - 1 GGGAAUUGAAAUUAAUUUGCUGCCAGAAUGCGGUGGGUGGAAAUAUGGGAGGAGGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAUUGACAUGGAACACACGCAAUCUGUCUCCUUCC ..............(((..((((......))))..)))........((((((((.......((((((((((...((......((((...)))).....))..)))))))))))))))))) ( -31.41) >DroEre_CAF1 158645 114 - 1 GGGAAUUGAAAUUAAUUUGCUGCCCAAAGGCGGUGGGUGGAAAUGUGGGC---GGGUUGGAAUAGAUUGUGCAUUGCAUAAUUACCAAUGGCACGGAACACACGCAAUCUUUCU---UCU (..(((((......(((..(((((....)))))..))).....((((..(---.((((((....((((((((...)))))))).)))))....).)..))))..)))).)..).---... ( -29.60) >consensus GGGAAUUGAAAUUAAUUUGCUGCCAGAAGGCGGUGGGUGGAAAUAUGGGA___GGAGUGGAAUAGAUUGUGCAUUGCAUAAUUGUCAAUGACAUGGAACACACGCAAUCUGUCUCCUUCC ..............(((..(((((....)))))..)))...............((((.(((.((((((((((((((((....)).)))))....(.....).)))))))))))).)))). (-26.12 = -27.62 + 1.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:56:26 2006