
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,622,875 – 16,622,972 |
| Length | 97 |
| Max. P | 0.967057 |

| Location | 16,622,875 – 16,622,972 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 81.34 |
| Mean single sequence MFE | -27.56 |
| Consensus MFE | -19.26 |
| Energy contribution | -19.86 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.39 |
| SVM RNA-class probability | 0.949682 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
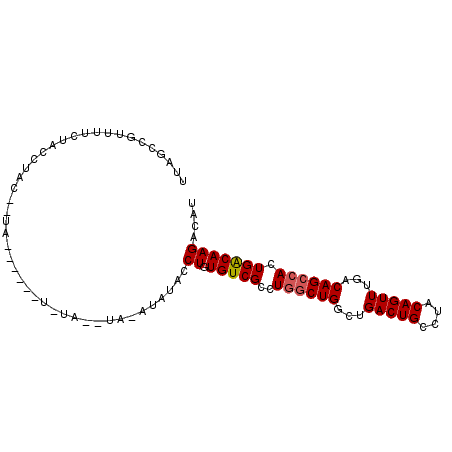
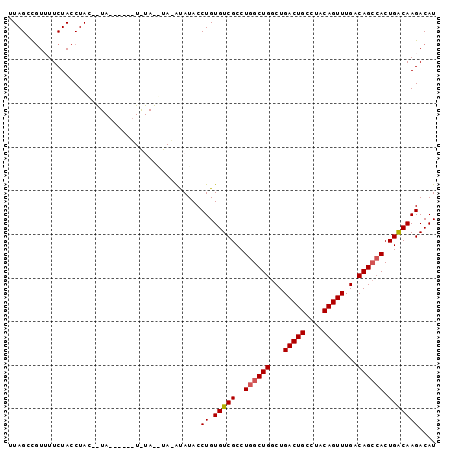
>2L_DroMel_CAF1 16622875 97 + 22407834 AUGUCUUGUCAGUGGCUGUCAAACUGUAGGCAGUCAGCCAGCCAGGCGACACAGGUAUGUAUAUGUAUAUAUAUGUAC-AUAGGUAGAAAACAGCUAA .(((.(((((..((((((.(..((((....))))..).))))))))))).)))..(((((((((((....))))))))-)))................ ( -29.90) >DroSec_CAF1 77810 79 + 1 AUGUCUUGUCAGUGGCUGUCAAACUGUAGGCAGUCAGCCAGCCAGGCGACACAGGUAUAU-------------------GUAGGUAGAAAACGUCUAA ((..((((((..((((((((........))))))))(((.....))))))).))..))..-------------------.....((((.....)))). ( -23.00) >DroSim_CAF1 76990 79 + 1 AUGUCUUGUCAGUGGCUGUCAAACUGUAGGCAGUCAGCCAGCCAGGCGACACAGGUAUAU-------------------GUAGGUAGAAAACGUCUAA ((..((((((..((((((((........))))))))(((.....))))))).))..))..-------------------.....((((.....)))). ( -23.00) >DroEre_CAF1 78960 97 + 1 AUGUCUUGCCAGUUCCUGUCAAACUGUAGGCAGUCAGCCAGCCAGGCGACACAGGUAUAUGUACAUAUACUCGUAUAUAGUAGGUAGAA-ACGGCUAA .......(((.(((.((..(..((((((.((.(((.(((.....))))))...(((((((....))))))).)).)))))).)..)).)-)))))... ( -28.10) >DroYak_CAF1 76847 90 + 1 AUGUCUUGUCAGUGGCUGUCAAACUGCAGGCAGUCAGCCAGCCAGGCGACACAGGUAUACCUACAUACA------UAU-GUAGGUAGAA-AGGGCUAA ..((((((((..((((((((........))))))))(((.....)))))).......(((((((((...------.))-)))))))...-)))))... ( -33.80) >consensus AUGUCUUGUCAGUGGCUGUCAAACUGUAGGCAGUCAGCCAGCCAGGCGACACAGGUAUAU_UA__UA_A______UA__GUAGGUAGAAAACGGCUAA ((..((((((..((((((((........))))))))(((.....))))))).))..))........................................ (-19.26 = -19.86 + 0.60)



| Location | 16,622,875 – 16,622,972 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 81.34 |
| Mean single sequence MFE | -25.24 |
| Consensus MFE | -18.14 |
| Energy contribution | -18.38 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.38 |
| Structure conservation index | 0.72 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.967057 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16622875 97 - 22407834 UUAGCUGUUUUCUACCUAU-GUACAUAUAUAUACAUAUACAUACCUGUGUCGCCUGGCUGGCUGACUGCCUACAGUUUGACAGCCACUGACAAGACAU .....((((((.....(((-(((........))))))...........((((..((((((...(((((....)))))...)))))).)))))))))). ( -24.30) >DroSec_CAF1 77810 79 - 1 UUAGACGUUUUCUACCUAC-------------------AUAUACCUGUGUCGCCUGGCUGGCUGACUGCCUACAGUUUGACAGCCACUGACAAGACAU ......(((((......((-------------------(......)))((((..((((((...(((((....)))))...)))))).))))))))).. ( -20.90) >DroSim_CAF1 76990 79 - 1 UUAGACGUUUUCUACCUAC-------------------AUAUACCUGUGUCGCCUGGCUGGCUGACUGCCUACAGUUUGACAGCCACUGACAAGACAU ......(((((......((-------------------(......)))((((..((((((...(((((....)))))...)))))).))))))))).. ( -20.90) >DroEre_CAF1 78960 97 - 1 UUAGCCGU-UUCUACCUACUAUAUACGAGUAUAUGUACAUAUACCUGUGUCGCCUGGCUGGCUGACUGCCUACAGUUUGACAGGAACUGGCAAGACAU ...(((((-((((...........(((.((((((....)))))).)))((((.(((...(((.....)))..)))..)))))))))).)))....... ( -28.10) >DroYak_CAF1 76847 90 - 1 UUAGCCCU-UUCUACCUAC-AUA------UGUAUGUAGGUAUACCUGUGUCGCCUGGCUGGCUGACUGCCUGCAGUUUGACAGCCACUGACAAGACAU ........-...(((((((-((.------...)))))))))...((.(((((..((((((...(((((....)))))...)))))).))))))).... ( -32.00) >consensus UUAGCCGUUUUCUACCUAC__UA______U_UA__UA_AUAUACCUGUGUCGCCUGGCUGGCUGACUGCCUACAGUUUGACAGCCACUGACAAGACAU ............................................((.(((((..((((((...(((((....)))))...)))))).))))))).... (-18.14 = -18.38 + 0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:55:16 2006