
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,589,105 – 16,589,413 |
| Length | 308 |
| Max. P | 0.993591 |

| Location | 16,589,105 – 16,589,210 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 97.33 |
| Mean single sequence MFE | -21.94 |
| Consensus MFE | -20.38 |
| Energy contribution | -20.76 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.984976 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16589105 105 + 22407834 ACGAUUUUUCCUUCUAGCCAAAUUGCUCCAAUUGCUAAUAAGGCUCACUCGAAAGCCACAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUU ..((((((......((((..(((((...)))))))))....((((....(....)....))))))))))...(((....(((((....))))).....))).... ( -22.20) >DroSec_CAF1 44405 105 + 1 CCGAUUUUUCCUUCUAGCCAAAUUGCUCCAAUUGCUAAUAAGGCUCACUCGAAAGCCACAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUU ..((((((......((((..(((((...)))))))))....((((....(....)....))))))))))...(((....(((((....))))).....))).... ( -22.20) >DroSim_CAF1 43214 105 + 1 CCGAUUUUUCCUUCUAGCCAAAUUGCUCCAAUUGCUAAUAAGGCUCACUCGAAAGCCACAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUU ..((((((......((((..(((((...)))))))))....((((....(....)....))))))))))...(((....(((((....))))).....))).... ( -22.20) >DroEre_CAF1 42812 105 + 1 CCGAUUUUUCCUUCUAGCCAAAUUGCUCCGAUUGCUAAUAAGGCUCACUCGAAAGCCACAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUU ..((((((......((((..(((((...)))))))))....((((....(....)....))))))))))...(((....(((((....))))).....))).... ( -21.90) >DroYak_CAF1 43023 105 + 1 CCGAUUUUUCCUUCUAGCCAAAUUGCUCCAAUUGCUAAUAAGGCUCACUUGAGAGCCGCAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGAGAUCAGACACUU ..((((((..((.(((((..(((((...)))))))))....(((((......)))))).))..))))))...(((..(..((((....))))..)...))).... ( -21.20) >consensus CCGAUUUUUCCUUCUAGCCAAAUUGCUCCAAUUGCUAAUAAGGCUCACUCGAAAGCCACAGCCAAAAUCCAAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUU ..((((((......((((..(((((...)))))))))....((((....(....)....))))))))))...(((....(((((....))))).....))).... (-20.38 = -20.76 + 0.38)


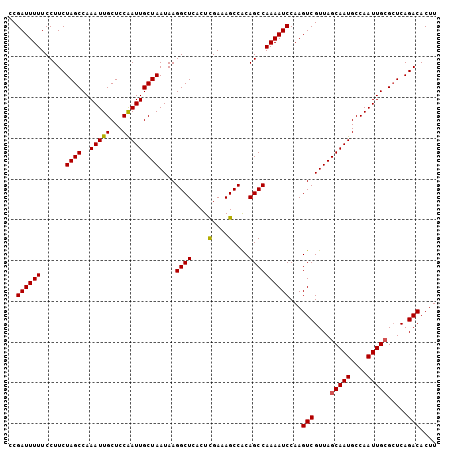
| Location | 16,589,175 – 16,589,276 |
|---|---|
| Length | 101 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 97.23 |
| Mean single sequence MFE | -24.20 |
| Consensus MFE | -22.60 |
| Energy contribution | -23.00 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.21 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.655471 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16589175 101 + 22407834 AAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUUUGGAACAAUCGGAAUCCCAGCCCGGAGUUGAAUCAGACGAUUUUCAUUUCGAUUUCCAUUACCAUU ..(((....(((((....))))).....)))....(((((..((((((((((......)))(..(((((....))))).).))))))))))))........ ( -23.00) >DroSec_CAF1 44475 101 + 1 AAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUUCGGAACAAUCGGAAUCCGAGCCCGGAGUUGAAUCAGACGAUUUUCAUUUCGAUUUCCAUUACCAUU ..(((....(((((....))))).....))).....((((..(((((((((((....))))(..(((((....))))).).)))))))))))......... ( -25.40) >DroSim_CAF1 43284 101 + 1 AAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUUCGGAACAAUCGGAAUCCGAGCCCGGAGUUGAAUCAGACGAUUUUCAUUUCGAUUUCCAUUACCAUU ..(((....(((((....))))).....))).....((((..(((((((((((....))))(..(((((....))))).).)))))))))))......... ( -25.40) >DroEre_CAF1 42882 101 + 1 AAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUUCGGAACAAUCGGAAUCCGAGCCCGGAGUUGAAUCAGACGUUAUUCAUUUCGAUUUCCAUUACCAUU ..(((....(((((....))))).....))).....((((..(((((((((((....))))..(((((........))))))))))))))))......... ( -25.10) >DroYak_CAF1 43093 101 + 1 AAGUCGUUAGCAAUGCCAAUUGAGAUCAGACACUUCGGAACAAUCGGAAUCCGAGCCCGGAGUUGAAUCAGACGAUUUUCAUUUCGAUUUCCAUUAUCAUU ((((((((..((((....)))).(((((...(((((((.....((((...))))..))))))))).))).)))))))).......(((.......)))... ( -22.10) >consensus AAGUCGUUAGCAAUGCCAAUUGCGCUCAGACACUUCGGAACAAUCGGAAUCCGAGCCCGGAGUUGAAUCAGACGAUUUUCAUUUCGAUUUCCAUUACCAUU ..(((....(((((....))))).....))).....((((..(((((((((((....))))(((......)))........)))))))))))......... (-22.60 = -23.00 + 0.40)

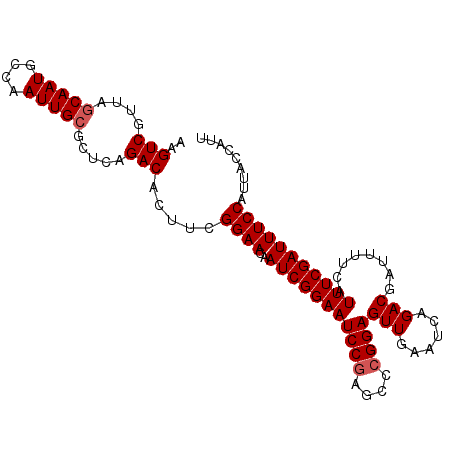

| Location | 16,589,210 – 16,589,316 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 97.74 |
| Mean single sequence MFE | -28.86 |
| Consensus MFE | -27.84 |
| Energy contribution | -27.88 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.96 |
| SVM decision value | 1.64 |
| SVM RNA-class probability | 0.969046 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16589210 106 - 22407834 CGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGGUAAUGGAAAUCGAAAUGAAAAUCGUCUGAUUCAACUCCGGGCUGGGAUUCCGAUUGUUCCA .(((((((((((.(.((..((((..(((((......))))).......((((...(((........))).....))))..))))..)).)..)))).))))))).. ( -26.40) >DroSec_CAF1 44510 106 - 1 CGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGGUAAUGGAAAUCGAAAUGAAAAUCGUCUGAUUCAACUCCGGGCUCGGAUUCCGAUUGUUCCG ((((((((((((.((((..((((..(((((......))))).......((((...(((........))).....))))..))))..))))..)))).)))).)))) ( -29.70) >DroSim_CAF1 43319 106 - 1 CGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGGUAAUGGAAAUCGAAAUGAAAAUCGUCUGAUUCAACUCCGGGCUCGGAUUCCGAUUGUUCCG ((((((((((((.((((..((((..(((((......))))).......((((...(((........))).....))))..))))..))))..)))).)))).)))) ( -29.70) >DroEre_CAF1 42917 106 - 1 CGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGGUAAUGGAAAUCGAAAUGAAUAACGUCUGAUUCAACUCCGGGCUCGGAUUCCGAUUGUUCCG ((((((((((((.((((..((((..(((((......)))))...................(((((........)))))..))))..))))..)))).)))).)))) ( -30.10) >DroYak_CAF1 43128 106 - 1 CGGACAAUGGAAAUGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGAUAAUGGAAAUCGAAAUGAAAAUCGUCUGAUUCAACUCCGGGCUCGGAUUCCGAUUGUUCCG ((((((((((((.((((..((((..(((((......))))).(((...((((...(((........)))))))..)))..))))..))))..)))).)))).)))) ( -28.40) >consensus CGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCUAAUGGUAAUGGAAAUCGAAAUGAAAAUCGUCUGAUUCAACUCCGGGCUCGGAUUCCGAUUGUUCCG .(((((((((((.((((..((((..(((((......))))).........(((.(((..(((.....))).))))))...))))..))))..)))).))))))).. (-27.84 = -27.88 + 0.04)

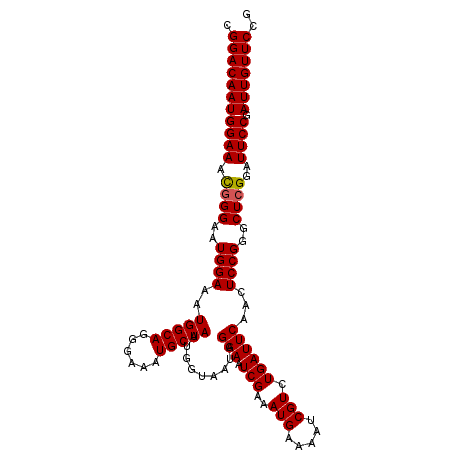

| Location | 16,589,276 – 16,589,388 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 95.21 |
| Mean single sequence MFE | -35.44 |
| Consensus MFE | -32.44 |
| Energy contribution | -32.24 |
| Covariance contribution | -0.20 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.40 |
| SVM RNA-class probability | 0.950392 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
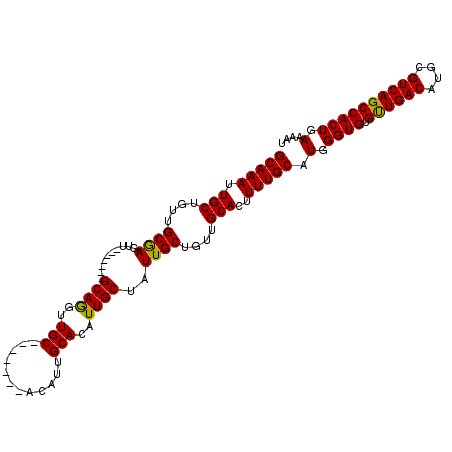
>2L_DroMel_CAF1 16589276 112 - 22407834 UUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCU ..((-------(..((((.((((.(((((.((((((((....))).(.((((.((((((....)))))))))).)...............))))).))))).)))))))).....))). ( -34.30) >DroSec_CAF1 44576 112 - 1 UUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCU ..((-------(..((((.((((.(((((.((((((((....))).(.((((.((((((....)))))))))).)...............))))).))))).)))))))).....))). ( -34.30) >DroSim_CAF1 43385 112 - 1 UUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCU ..((-------(..((((.((((.(((((.((((((((....))).(.((((.((((((....)))))))))).)...............))))).))))).)))))))).....))). ( -34.30) >DroEre_CAF1 42983 119 - 1 UUGCACAUUGCACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGCUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCU ..(((..((((.((((...((((.(((((((((.((((....))))(.((((.((((((....)))))))))).)....))).))))(....))).))))..)))))))).....))). ( -37.40) >DroYak_CAF1 43194 119 - 1 UUGCACAUUGCACAUUGCACAUUGCUCUUGCUGUUGCACUUUUGCAUGGGUGUGCUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAAUGGGAAUGGAAAUGGCAGGGAAAUGCU ((((.((((...((((.(.((((.((.((((((.((((....))))(.((((.((((((....)))))))))).)....))).))).)).))))).))))..))))))))......... ( -36.90) >consensus UUGC_______ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAUCGGACAAUGGAAACGGGAAUGGAAAUGGCAGGGAAAUGCU ..............((((.((((.(((((.((((((((....))).(.((((.((((((....)))))))))).)...............))))).))))).))))))))......... (-32.44 = -32.24 + -0.20)



| Location | 16,589,316 – 16,589,413 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 91.02 |
| Mean single sequence MFE | -37.78 |
| Consensus MFE | -28.90 |
| Energy contribution | -28.34 |
| Covariance contribution | -0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.53 |
| Structure conservation index | 0.76 |
| SVM decision value | 2.41 |
| SVM RNA-class probability | 0.993591 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 16589316 97 - 22407834 GCAAAUUGCUGUUGCGAGUU-------GCAGGUUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAU (((((.(((....((((...-------((((..(((-------.....)))..))))..))))....)))..))))).(.((((.((((((....)))))))))).).... ( -33.90) >DroSec_CAF1 44616 97 - 1 GCAAAUUGCUGUUGCGAGUU-------GCAGGUUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAU (((((.(((....((((...-------((((..(((-------.....)))..))))..))))....)))..))))).(.((((.((((((....)))))))))).).... ( -33.90) >DroSim_CAF1 43425 97 - 1 GCAAAUUGCUGUUGCGAGUU-------GCAGGUUGC-------ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAU (((((.(((....((((...-------((((..(((-------.....)))..))))..))))....)))..))))).(.((((.((((((....)))))))))).).... ( -33.90) >DroEre_CAF1 43023 111 - 1 GCAAAUUGCAGUUGCAAGUUGCAAGUUGCAAGUUGCACAUUGCACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGCUGACAUGCGUCAGCCACUGAAAAU (((((.(((((..((((...((((..(((((..(((.....)))..)))))..))))..))))..)))))..))))).(.((((.((((((....)))))))))).).... ( -46.20) >DroYak_CAF1 43234 104 - 1 GCAAAUUGCAGUUGCGAGUU-------GCAGGUUGCACAUUGCACAUUGCACAUUGCUCUUGCUGUUGCACUUUUGCAUGGGUGUGCUGACAUGCGUCAGCCACUGAAAAU (((((.(((((..(((((..-------((((..((((..........))))..)))).)))))..)))))..))))).(.((((.((((((....)))))))))).).... ( -41.00) >consensus GCAAAUUGCUGUUGCGAGUU_______GCAGGUUGC_______ACAUUGCACAUUGCUAUUGCUGUUGCACUUUUGCAUGGGUGUGUUGACAUGCGUCAGCCACUGAAAAU (((((.(((....((((..........((((..(((............)))..))))..))))....)))..))))).(.((((.((((((....)))))))))).).... (-28.90 = -28.34 + -0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:54:59 2006