
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,781,305 – 1,781,405 |
| Length | 100 |
| Max. P | 0.791687 |

| Location | 1,781,305 – 1,781,405 |
|---|---|
| Length | 100 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 74.85 |
| Mean single sequence MFE | -31.85 |
| Consensus MFE | -15.64 |
| Energy contribution | -18.07 |
| Covariance contribution | 2.44 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.49 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.791687 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1781305 100 + 22407834 AGUGCAAAUAUUUUCG-------------ACCGAUUUAGCGCUUCAUUUGCCUGGGUGGCCGUGACCGGGAGGGGCGUGGCAGUUUGUUAUUCUUUGAUAUGCUAAAUUGGUG ................-------------((((((((((((..(((...((....)).((((((.((....))..))))))..............)))..)))))))))))). ( -29.80) >DroPse_CAF1 45113 108 + 1 UGUGCAAAUAUUUUCCCACUUCGUUUUGCCCCGUUUUUGCACUUCAUUUGUGUUUGAAGGGG-----AAGAGGGGAGUGGCUGUUUGCUUUUCUUUGAUAUGCGAAAUUUAUG ...(((((((.....(((((((.((((.((((.(((..((((.......))))..)))))))-----.)))).))))))).)))))))......................... ( -33.30) >DroSec_CAF1 38164 100 + 1 AGUGCAAAUAUUUUCG-------------ACCGAUUUUGCACUUCAUUUGCCUGGGAGGGCGUGUCCGGGAGGGACGUGGCAGUUUGUUUUUCUUUGAUGUGCUAAAUUGGUG ................-------------((((((((.((((.......((((....))))..(((..(((((((((........)))))))))..))))))).)))))))). ( -31.00) >DroPer_CAF1 43716 108 + 1 UGUGCAAAUAUUUUCCCACUUCGUUUUGCCCCGUUUUUGCACUUCAUUUGUGUUUGAAGGGG-----AAGAGGGGAGUGGCUGUUUGCUUUUCUUUGAUAUGCGAAAUUUAUG ...(((((((.....(((((((.((((.((((.(((..((((.......))))..)))))))-----.)))).))))))).)))))))......................... ( -33.30) >consensus AGUGCAAAUAUUUUCC_____________ACCGAUUUUGCACUUCAUUUGCCUGGGAAGGCG_____AAGAGGGGAGUGGCAGUUUGCUUUUCUUUGAUAUGCGAAAUUGAUG .............................(((((((((((((((((........))))).........(((((((((..(....)..)))))))))....)))))))))))). (-15.64 = -18.07 + 2.44)

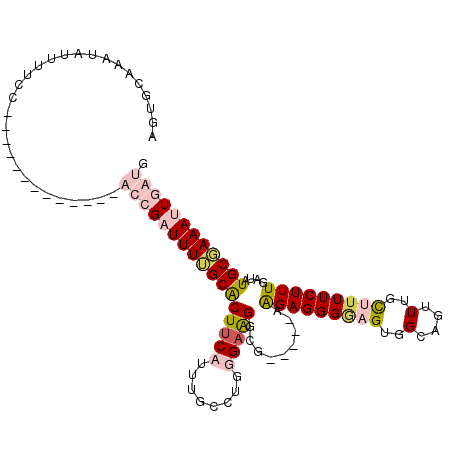
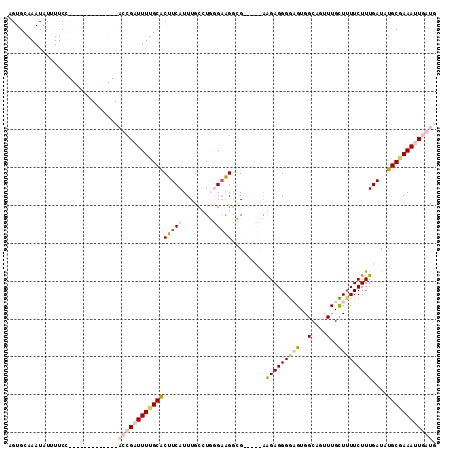
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:40:15 2006