| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,397,271 – 16,397,368 |
| Length | 97 |
| Max. P | 0.991546 |

| Location | 16,397,271 – 16,397,368 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.40 |
| Mean single sequence MFE | -34.33 |
| Consensus MFE | -12.90 |
| Energy contribution | -14.18 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.38 |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.922626 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
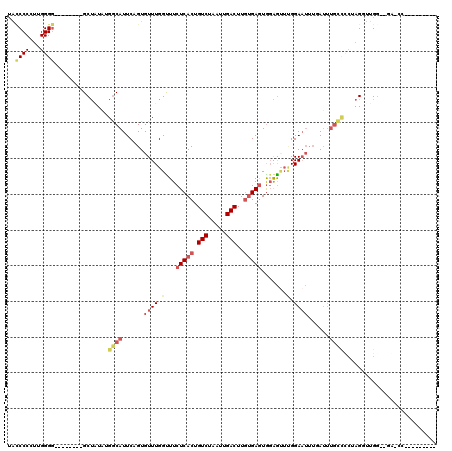
>2L_DroMel_CAF1 16397271 97 + 22407834 UAUCCCCUUGGGG--------GCUAUAUGGCAUUCUGUGUUUGGUUUCUCACUGUCUAAUUGACUUGUGAGUGGAGCUUGGAAUUUUAUU------UAGUUUGGGGGGGCC--------- ..((((((..(..--------((((.((((.((((((.((((.(...(((((.(((.....)))..)))))).)))).)))))).)))).------)))))..))))))..--------- ( -33.00) >DroSec_CAF1 21734 101 + 1 UACCCCCUUGGGG--------GCUAUAUGGCAUUCUGUGUUUGGUUUCUCACUGUCUAAUUGACUUGUGAGUGGAGUUUGGAAUUUGAAAUGCCCCUAGGUUGG--GGGCC--------- ..(((((((((((--------((........((((((..(((.(...(((((.(((.....)))..)))))).)))..)))))).......)))))))))..))--))...--------- ( -36.86) >DroSim_CAF1 20489 101 + 1 UACCCCCUUGGGG--------GCUUUAUGGCAUUCUGUGUUUGGUUUCUCACUGUCUAAUUGACUUGUGAGUAGAGUUUGGAAUUUUAUUUGCCCCUAGGUUGG--GGCCC--------- ..(((((((((((--------((...((((.((((((..(((.....(((((.(((.....)))..)))))..)))..)))))).))))..)))))))))..))--))...--------- ( -39.40) >DroEre_CAF1 18100 98 + 1 UAACCCCUGGGG---------GCUGUGUGGCAUUCAGUGUUUGGCUUCUCACUGUCUAAUUGACUUGUGAGUGGAGUUGGGAAUGUGAUUUGCCUC-AGUUUGG--GA-AA--------- ....(((..(((---------((...((.((((((.....(..(((((((((.(((.....)))..)))))..))))..))))))).))..)))))-.....))--).-..--------- ( -35.20) >DroYak_CAF1 22365 107 + 1 UACCCCCUUGGGGGCUAUUUGGCUAUGUGGCAUUCAGUGUUUGGUUUCUCACUGUCUAAUUGACUUGGGAGUGCAGUUCGGAAUUUGAUUUGCCUC-AGAUUGG--GA-AA--------- ..((((....))))...((..(...((.((((.((((..(((((.((((((..(((.....))).))))))......)))))..))))..)))).)-)..)..)--).-..--------- ( -28.50) >DroAna_CAF1 17141 111 + 1 GGCCCCCCAGGGG--------GCAAUAUGCCAUUCAGUGUUUGGUUUCUCACUGUCUAAUUGACUUCUGAAUGGGAAAUGGACAUGCAUUUGGUCCAAGGAUGAGGGA-GGCUUUAAAUU (((((((((..((--------((...((((......((((.(.((((((((..(((.....))).......)))))))).)))))))))...)))).....)).))).-))))....... ( -33.00) >consensus UACCCCCUUGGGG________GCUAUAUGGCAUUCAGUGUUUGGUUUCUCACUGUCUAAUUGACUUGUGAGUGGAGUUUGGAAUUUGAUUUGCCCCUAGGUUGG__GA_CC_________ ..((((...))))...............((((......((((.(...(((((.(((.....)))..))))).......).))))......)))).......................... (-12.90 = -14.18 + 1.28)


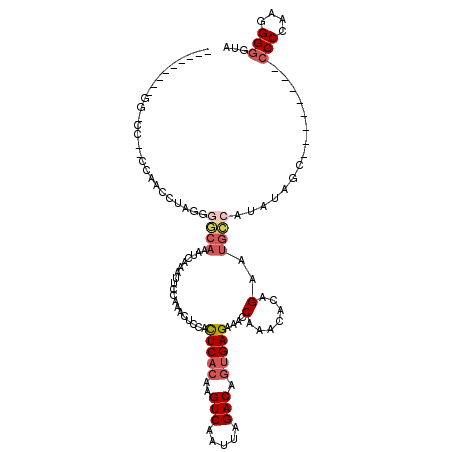
| Location | 16,397,271 – 16,397,368 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 75.40 |
| Mean single sequence MFE | -25.72 |
| Consensus MFE | -10.61 |
| Energy contribution | -11.58 |
| Covariance contribution | 0.97 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.98 |
| Structure conservation index | 0.41 |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.991546 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
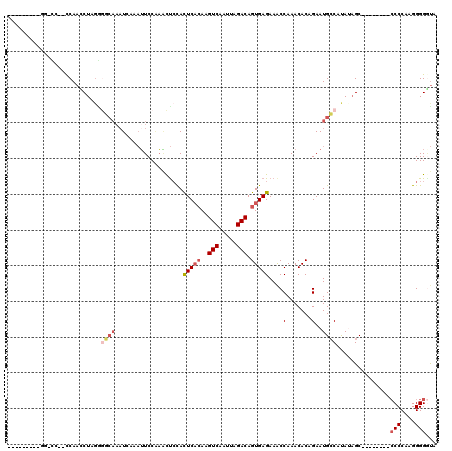
>2L_DroMel_CAF1 16397271 97 - 22407834 ---------GGCCCCCCCAAACUA------AAUAAAAUUCCAAGCUCCACUCACAAGUCAAUUAGACAGUGAGAAACCAAACACAGAAUGCCAUAUAGC--------CCCCAAGGGGAUA ---------.....((((......------...................(((((..(((.....))).)))))................((......))--------......))))... ( -16.90) >DroSec_CAF1 21734 101 - 1 ---------GGCCC--CCAACCUAGGGGCAUUUCAAAUUCCAAACUCCACUCACAAGUCAAUUAGACAGUGAGAAACCAAACACAGAAUGCCAUAUAGC--------CCCCAAGGGGGUA ---------(((((--((......)))).....................(((((..(((.....))).)))))................))).....((--------(((....))))). ( -30.00) >DroSim_CAF1 20489 101 - 1 ---------GGGCC--CCAACCUAGGGGCAAAUAAAAUUCCAAACUCUACUCACAAGUCAAUUAGACAGUGAGAAACCAAACACAGAAUGCCAUAAAGC--------CCCCAAGGGGGUA ---------(((((--((......)))))....................(((((..(((.....))).)))))...))...................((--------(((....))))). ( -32.00) >DroEre_CAF1 18100 98 - 1 ---------UU-UC--CCAAACU-GAGGCAAAUCACAUUCCCAACUCCACUCACAAGUCAAUUAGACAGUGAGAAGCCAAACACUGAAUGCCACACAGC---------CCCCAGGGGUUA ---------..-..--.......-..((((...................(((((..(((.....))).)))))....((.....))..))))....(((---------((....))))). ( -21.10) >DroYak_CAF1 22365 107 - 1 ---------UU-UC--CCAAUCU-GAGGCAAAUCAAAUUCCGAACUGCACUCCCAAGUCAAUUAGACAGUGAGAAACCAAACACUGAAUGCCACAUAGCCAAAUAGCCCCCAAGGGGGUA ---------..-..--.......-..((((...........((..((......))..)).......(((((..........)))))..)))).............(((((....))))). ( -22.00) >DroAna_CAF1 17141 111 - 1 AAUUUAAAGCC-UCCCUCAUCCUUGGACCAAAUGCAUGUCCAUUUCCCAUUCAGAAGUCAAUUAGACAGUGAGAAACCAAACACUGAAUGGCAUAUUGC--------CCCCUGGGGGGCC ........(((-((((.......(((((.........)))))....((((((((..(((.....))).((.....))......))))))))........--------.....))))))). ( -32.30) >consensus _________GG_CC__CCAACCUAGGGGCAAAUCAAAUUCCAAACUCCACUCACAAGUCAAUUAGACAGUGAGAAACCAAACACAGAAUGCCAUAUAGC________CCCCAAGGGGGUA ..........................((((...................(((((..(((.....))).)))))....(.......)..))))...............(((....)))... (-10.61 = -11.58 + 0.97)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:42 2006