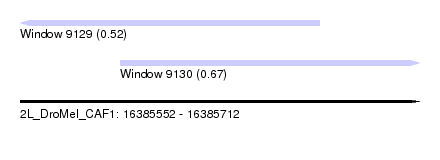
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,385,552 – 16,385,712 |
| Length | 160 |
| Max. P | 0.667891 |

| Location | 16,385,552 – 16,385,672 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 90.75 |
| Mean single sequence MFE | -29.40 |
| Consensus MFE | -22.37 |
| Energy contribution | -22.77 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.524309 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
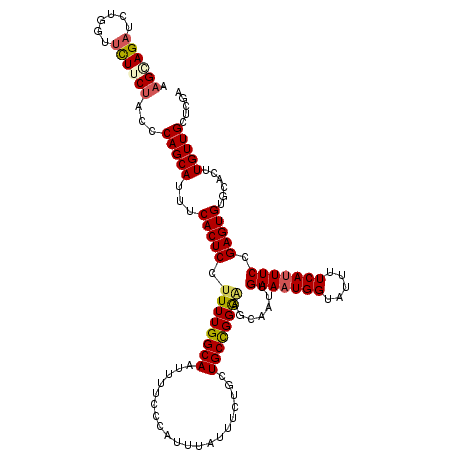
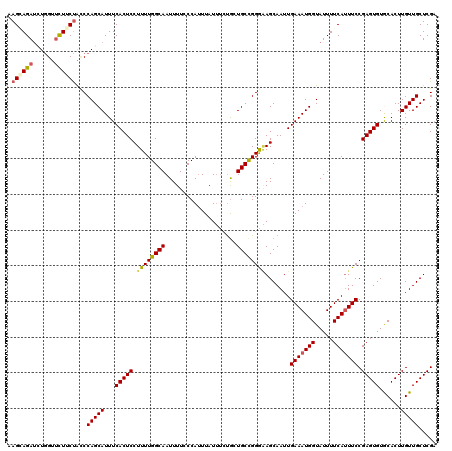
>2L_DroMel_CAF1 16385552 120 - 22407834 AAGGAGAUAUGGUUCUUCUAUCCAGCAUUUCACUCCUUUUGGCAAUUUUCCCAUUUAUUUCUGCUGCUGGGAAGCAAUUGAAAUGGUAUUUUCAUUUCCGAGUGUGCACUUGUUGCUCGA .((((((......)))))).(((((((............(((........)))...........))))))).(((((..(((((((.....)))))))(((((....))))))))))... ( -29.40) >DroSec_CAF1 9082 120 - 1 AAGGAGAUCUGGUUCUUCUAACCAGCAUUUCACUCCUUUUGGCAAUUUUCCCAUUUAUUUCUGCUGCCGGAGAGCAAUUGAAAUGGUAUUUUCAUUUCCGAGUGUGCACUUGUUGCUCGA ...(((((((((((.....)))))).)))))......(((((((....................)))))))((((((..(((((((.....)))))))(((((....))))))))))).. ( -33.65) >DroSim_CAF1 7196 120 - 1 AAGAAGAACUGGUUCUUCUACCCAGCAUUUCACUCCUUUUGGCAAUUUUCCCAUUUAUUUCUGCUGCCGGAAAGCAAUUGAAAUGGUAUUUUCAUUUCCGAGUGUGCACUUGUUGCUCGA .(((((((....)))))))...(((((...(((((.((((((((....................)))))))).......(((((((.....))))))).)))))......)))))..... ( -29.05) >DroEre_CAF1 7243 120 - 1 CAGCAGAUUUUGCAUUUCCGCCCAGCAUUUCACUCCUUUUGGCAAUUUUCCCAUUUAUGGCCGCUGCCGGGUAGCAAUUGAAAUGGUAUUUUCAGUUCCGAGUGUACACUUGUUGCUCGC ((((.((...(((...........)))..)).........(((................))))))).((((((((((((((((......))))))))..((((....)))))))))))). ( -27.49) >DroYak_CAF1 10429 120 - 1 AAGCAGAUUUGGUUCUUCUACCCAGCAUUUCACUCCUUUUGGCAAUUUUCCCACUUAUUUAUACUGCCGGGGAGCAAUUGAAAUGGUAUUUUCAUUUCCGAGUGUACACUUGUUGCUCGA .((.(((......))).)).(((.(((............(((........)))...........))).)))((((((..(((((((.....)))))))(((((....))))))))))).. ( -27.40) >consensus AAGCAGAUCUGGUUCUUCUACCCAGCAUUUCACUCCUUUUGGCAAUUUUCCCAUUUAUUUCUGCUGCCGGGAAGCAAUUGAAAUGGUAUUUUCAUUUCCGAGUGUGCACUUGUUGCUCGA .((((((......))))))...(((((...(((((.((((((((....................)))))))).......(((((((.....))))))).)))))......)))))..... (-22.37 = -22.77 + 0.40)

| Location | 16,385,592 – 16,385,712 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.56 |
| Mean single sequence MFE | -30.10 |
| Consensus MFE | -18.17 |
| Energy contribution | -18.53 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.667891 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
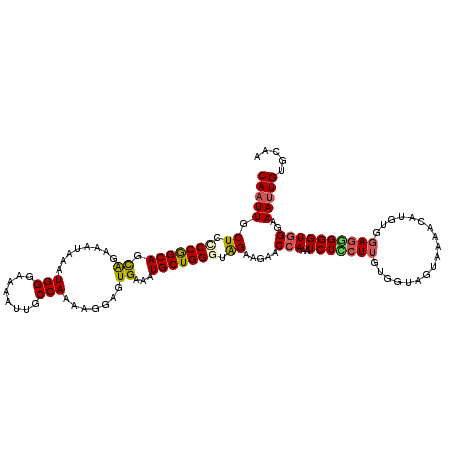
>2L_DroMel_CAF1 16385592 120 + 22407834 CAAUUGCUUCCCAGCAGCAGAAAUAAAUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGAUAGAAGAACCAUAUCUCCUUGUGGUAGUAAAACAUGUGGAGGGGGUGGGAAAUUGUGCGA (((((.((((((((((.((........(((........))).......))...))))))...))))..((((.(((((.((((.........)))).)))))..))))..)))))..... ( -29.36) >DroSec_CAF1 9122 120 + 1 CAAUUGCUCUCCGGCAGCAGAAAUAAAUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGUUAGAAGAACCAGAUCUCCUUGUGGUAGUAAAACAUGUGGAGGGGGUGGGAAAUUGUGCAA ...(..((((((.(((.((........)).....(((((((.((((((.......((((((.....))))))..)))))).)))))))......))).).)))))..)............ ( -34.90) >DroSim_CAF1 7236 120 + 1 CAAUUGCUUUCCGGCAGCAGAAAUAAAUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGGUAGAAGAACCAGUUCUUCUUGUGGUAGUAAAACAUGUGGAGGGGGUGGGAAAUUGUGCAA ...(((((((((.((..(........(((.....(((((((.((((((......(((((.........))))).)))))).)))))))....)))......)..)).)))))....)))) ( -30.54) >DroEre_CAF1 7283 103 + 1 CAAUUGCUACCCGGCAGCGGCCAUAAAUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGGCGGAAAUGCAAAAUCUGCUGGU---------------GGAGUGGGUGGGAAAAUGU--AG ......(((((((.(((((((((....))).....(((((...((.(......).)).)))))...........)))))).)---------------)).))))............--.. ( -27.40) >DroYak_CAF1 10469 118 + 1 CAAUUGCUCCCCGGCAGUAUAAAUAAGUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGGUAGAAGAACCAAAUCUGCUUGUGGUAGUAGAAAAUGUGGAAGGGGUGGGAAAUUGU--AG ...(..((((((((((.(((.......(((........)))......)))...)))))).........(((..((((((......)))))).....)))..))))..)........--.. ( -28.32) >consensus CAAUUGCUCCCCGGCAGCAGAAAUAAAUGGGAAAAUUGCCAAAAGGAGUGAAAUGCUGGGUAGAAGAACCAAAUCUCCUUGUGGUAGUAAAACAUGUGGAGGGGGUGGGAAAUUGUGCAA (((((.((.(((((((.((........(((........))).......))...))))))).)).....(((..(((((((..................))))))))))..)))))..... (-18.17 = -18.53 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:52:08 2006