
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 16,268,602 – 16,268,721 |
| Length | 119 |
| Max. P | 0.979842 |

| Location | 16,268,602 – 16,268,721 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 93.22 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -20.30 |
| Energy contribution | -20.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.40 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
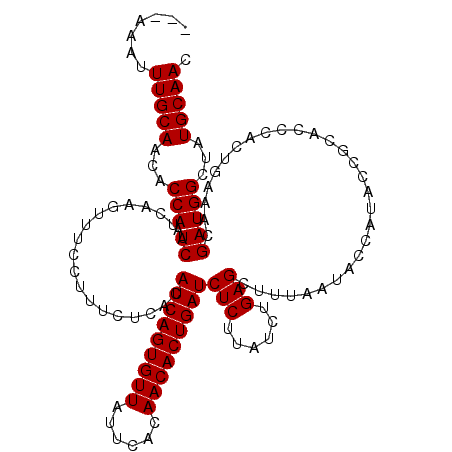
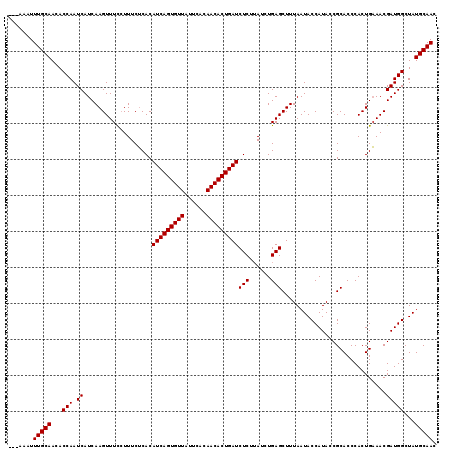
>2L_DroMel_CAF1 16268602 119 + 22407834 AAUACAUUUGCAAUACCAAUCAUCAAGUUUUCUUUCACACAUCAGUGUUGUUCACAACACUGAUCUCUAAUCUGAGCUUUAAUACCAUAACGCACCCACUGUAACGAUGGCUAUGCAAC .......(((((............((((((..........((((((((((....)))))))))).........)))))).....((((...(((.....)))....))))...))))). ( -24.71) >DroSec_CAF1 14393 116 + 1 ---AAAUUUGCAACACCAAUCAUCAAGUUUCCUUUCUCACAUCAGUGUUAUUCACAACACUGAUCUCUUAUCUGAGCUUUAAUACCAUACCGCACCCACUGAAACGAUGGCUAUGCAAC ---....(((((...(((.((.....(((((.........(((((((((......)))))))))(((......)))........................))))))))))...))))). ( -21.20) >DroSim_CAF1 10045 115 + 1 ---AAAUUUGCAACACCAAUCAUCAAGUUUC-UUUCUCACAUCAGUGUUAUUCACAACACUGAUCUCUUAUCUGAGCUUUAAUACCAUACCGCACCCACUGAAACGAUGGCUAUGCAAC ---....(((((...(((.((.....(((((-........(((((((((......)))))))))(((......)))........................))))))))))...))))). ( -21.60) >consensus ___AAAUUUGCAACACCAAUCAUCAAGUUUCCUUUCUCACAUCAGUGUUAUUCACAACACUGAUCUCUUAUCUGAGCUUUAAUACCAUACCGCACCCACUGAAACGAUGGCUAUGCAAC .......(((((...(((.((...................(((((((((......)))))))))(((......))).............................)))))...))))). (-20.30 = -20.30 + 0.00)

| Location | 16,268,602 – 16,268,721 |
|---|---|
| Length | 119 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 93.22 |
| Mean single sequence MFE | -34.07 |
| Consensus MFE | -31.22 |
| Energy contribution | -31.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.03 |
| Mean z-score | -3.13 |
| Structure conservation index | 0.92 |
| SVM decision value | 1.85 |
| SVM RNA-class probability | 0.979842 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
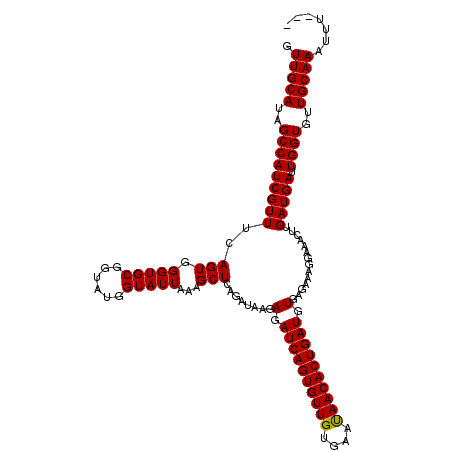
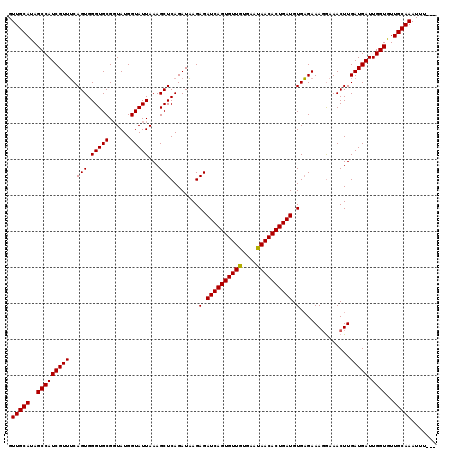
>2L_DroMel_CAF1 16268602 119 - 22407834 GUUGCAUAGCCAUCGUUACAGUGGGUGCGUUAUGGUAUUAAAGCUCAGAUUAGAGAUCAGUGUUGUGAACAACACUGAUGUGUGAAAGAAAACUUGAUGAUUGGUAUUGCAAAUGUAUU .(((((..((((((((((.(((((((...(((......))).))))...(((.(.((((((((((....)))))))))).).)))......))))))))).))))..)))))....... ( -38.80) >DroSec_CAF1 14393 116 - 1 GUUGCAUAGCCAUCGUUUCAGUGGGUGCGGUAUGGUAUUAAAGCUCAGAUAAGAGAUCAGUGUUGUGAAUAACACUGAUGUGAGAAAGGAAACUUGAUGAUUGGUGUUGCAAAUUU--- .(((((..((((((((((((((.(((((......)))))...)))........(.((((((((((....)))))))))).)..))(((....)))))))).))))..)))))....--- ( -33.00) >DroSim_CAF1 10045 115 - 1 GUUGCAUAGCCAUCGUUUCAGUGGGUGCGGUAUGGUAUUAAAGCUCAGAUAAGAGAUCAGUGUUGUGAAUAACACUGAUGUGAGAAA-GAAACUUGAUGAUUGGUGUUGCAAAUUU--- .(((((..(((((((((..(((.(((((......)))))....(((......)))((((((((((....))))))))))........-...))).))))).))))..)))))....--- ( -30.40) >consensus GUUGCAUAGCCAUCGUUUCAGUGGGUGCGGUAUGGUAUUAAAGCUCAGAUAAGAGAUCAGUGUUGUGAAUAACACUGAUGUGAGAAAGGAAACUUGAUGAUUGGUGUUGCAAAUUU___ .(((((..(((((((((..(((.(((((......)))))...)))........(.((((((((((....)))))))))).)..............))))).))))..)))))....... (-31.22 = -31.00 + -0.22)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:50:57 2006