
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,972,002 – 15,972,112 |
| Length | 110 |
| Max. P | 0.900086 |

| Location | 15,972,002 – 15,972,112 |
|---|---|
| Length | 110 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.33 |
| Mean single sequence MFE | -42.63 |
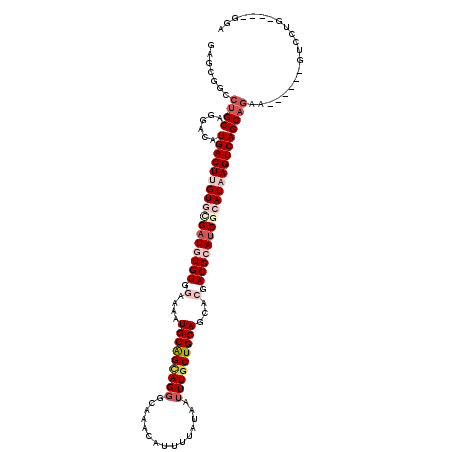
| Consensus MFE | -31.73 |
| Energy contribution | -33.84 |
| Covariance contribution | 2.11 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.44 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.900086 |
| Prediction | RNA |
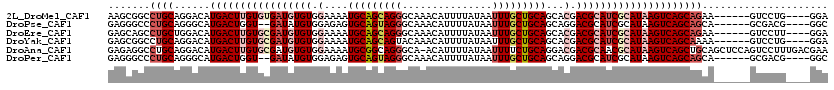
Download alignment: ClustalW | MAF
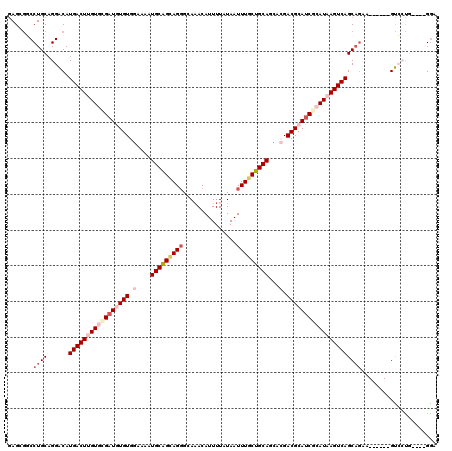
>2L_DroMel_CAF1 15972002 110 - 22407834 AAGCGGCCUGCAGGACAUGACUUGUGUGAUGUGUGGAAAAUGCAGCAGGGCAAACAUUUUAUAAUUUGCUGCAGCACGACGCAUCGCAUAAGUCAGCAGAA------GUCCUG----GGA ......((..((((((.(((((((((((((((((.(....(((((((((...............)))))))))...).)))))))))))))))))......------))))))----)). ( -48.36) >DroPse_CAF1 152480 108 - 1 GAGGGCCCUGCAGGGCAUGACUGGU--GAUAUGUGGAGAGUGCAGUAGGGCAAACAUUUUAUAAUUUGCUGCAGCAGGACGCAUCGCAUAAGUCAGCAGCA------GCGACG----GGC ....(((((((...((.(((((.((--(..(((((.....(((((((((...............)))))))))......)))))..))).)))))))....------)))..)----))) ( -33.46) >DroEre_CAF1 134420 110 - 1 GAGCAGCCUGCUGGACAUGACUUGUGCGAUGUGUGGAAAAUGCAGCAGGGCAAACAUUUUAUAAUUUGCUGCAGCACGACGCAUCGCAUAAGUCAGCAGAA------GUCCUU----GGA .(((.....)))((((.(((((((((((((((((.(....(((((((((...............)))))))))...).)))))))))))))))))......------))))..----... ( -47.76) >DroYak_CAF1 131938 110 - 1 GAGCGGCCUGCAGGACAUGACUUGUGCGAUGUGUGGAAAAUGCAGCAGUACAAACAUUUUAUAAUUUGCUGCAGCACGACGCAUCGCAUAAGUCAGCAAAA------GUCCUG----GGA ......((..((((((.(((((((((((((((((.(....((((((((.................))))))))...).)))))))))))))))))......------))))))----)). ( -49.83) >DroAna_CAF1 83531 119 - 1 GAGAGGCCUGCAGGACAUGACUUGUGCGAUGUGUGGAAAAUGCGGCAGGGCA-ACAUUUUAUAAUUUUCUGCAGGACGACGCAACGCAUAAGUCAGCUGCAGCUCCAGUCCUUUGACGAA ..(..((.(((((....(((((((((((.(((((.(....(((((...(...-.).............)))))...).))))).))))))))))).)))))))..).(((....)))... ( -42.89) >DroPer_CAF1 181191 108 - 1 GAGGGCCCUGCAGGGCAUGACUGGU--GAUAUGUGGAGAGUGCAGUAGGGCAAACAUUUUAUAAUUUGCUGCAGCAGGACGCAUCGCAUAAGUCAGCAGCA------GCGACG----GGC ....(((((((...((.(((((.((--(..(((((.....(((((((((...............)))))))))......)))))..))).)))))))....------)))..)----))) ( -33.46) >consensus GAGCGGCCUGCAGGACAUGACUUGUGCGAUGUGUGGAAAAUGCAGCAGGGCAAACAUUUUAUAAUUUGCUGCAGCACGACGCAUCGCAUAAGUCAGCAGAA______GUCCUG____GGA .......((((......(((((((((((((((((.(....(((((((((...............)))))))))...).)))))))))))))))))))))..................... (-31.73 = -33.84 + 2.11)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:49:02 2006