
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,839,093 – 15,839,283 |
| Length | 190 |
| Max. P | 0.702438 |

| Location | 15,839,093 – 15,839,213 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.17 |
| Mean single sequence MFE | -22.81 |
| Consensus MFE | -18.14 |
| Energy contribution | -18.37 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.80 |
| SVM decision value | -0.07 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
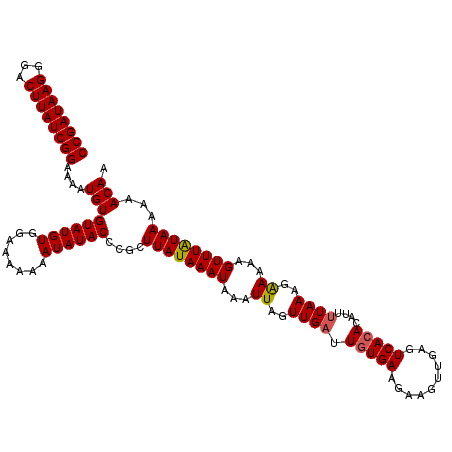
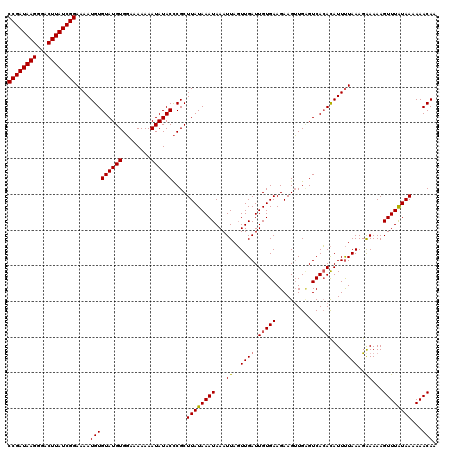
>2L_DroMel_CAF1 15839093 120 - 22407834 CCGAUAAGGGACUUAUCGGAAAAUGUGUAUGUGUAAAAAAAUAUACUCGCUUAUAAAUAAAUUAUUUGAUUGUGAAGAAGAUGUGUCAGAUAUUCUAAAGAAAAAGUUUGUAAAAAACAA ((((((((...)))))))).....(.((((((........)))))).)..((((((((....((((((((..............))))))))(((....)))...))))))))....... ( -22.24) >DroSec_CAF1 11105 120 - 1 CCGAUAAGGGACUUAUCGGAAAAUGUGUAUGUGGAAAAAAAUAUACCCGCUUAUAAAUAAAUUAGUUGAUUGUGAAGAAGUUGAGUCACACAUUUUAAAGAAAAAGUUUAUAAAAAACAA ((((((((...))))))))(((((((((..((((............))))((((((.(((.....))).))))))............))))))))).........((((.....)))).. ( -22.70) >DroSim_CAF1 11079 120 - 1 CCGAUAAGGGACUUAUCGGAAAAUGUGUAUGUGGAAAAAAAUAUACCCGCUUAUAAAUAAAUUAGUUGAUUGUGAAGAAGUUGCGUCACACAUUUUAAAGGAAAAGUUUAUAAAAAACAA ((((((((...))))))))(((((((((..((((............)))).................(((.(..(.....)..))))))))))))).........((((.....)))).. ( -23.50) >consensus CCGAUAAGGGACUUAUCGGAAAAUGUGUAUGUGGAAAAAAAUAUACCCGCUUAUAAAUAAAUUAGUUGAUUGUGAAGAAGUUGAGUCACACAUUUUAAAGAAAAAGUUUAUAAAAAACAA ((((((((...))))))))....(((((((((........))))))....((((((((...((..((((.(((((..........)))))....))))..))...))))))))...))). (-18.14 = -18.37 + 0.23)

| Location | 15,839,133 – 15,839,253 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 98.33 |
| Mean single sequence MFE | -17.47 |
| Consensus MFE | -17.22 |
| Energy contribution | -17.00 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.43 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.06 |
| SVM RNA-class probability | 0.562875 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
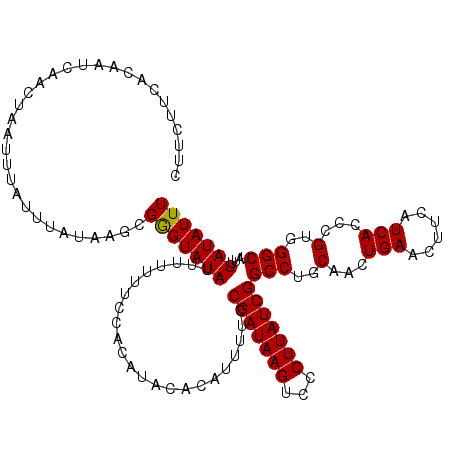
>2L_DroMel_CAF1 15839133 120 + 22407834 CUUCUUCACAAUCAAAUAAUUUAUUUAUAAGCGAGUAUAUUUUUUUACACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUU ................................(((((((.......................(((((((...)))))))(((..(...(((......)))...)..)))....))))))) ( -17.60) >DroSec_CAF1 11145 120 + 1 CUUCUUCACAAUCAACUAAUUUAUUUAUAAGCGGGUAUAUUUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUU ..............................((((((.........................((((((((...)))))))).........((......))))))))............... ( -17.40) >DroSim_CAF1 11119 120 + 1 CUUCUUCACAAUCAACUAAUUUAUUUAUAAGCGGGUAUAUUUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUU ..............................((((((.........................((((((((...)))))))).........((......))))))))............... ( -17.40) >consensus CUUCUUCACAAUCAACUAAUUUAUUUAUAAGCGGGUAUAUUUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUU ................................(((((((.......................(((((((...)))))))(((..(...(((......)))...)..)))....))))))) (-17.22 = -17.00 + -0.22)


| Location | 15,839,173 – 15,839,283 |
|---|---|
| Length | 110 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.89 |
| Mean single sequence MFE | -23.47 |
| Consensus MFE | -21.33 |
| Energy contribution | -21.67 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.702438 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15839173 110 + 22407834 UUUUUUACACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUUAUUACUAUUUAUUGGCUUAUGAGUCUGGCA---------- ......................(((((((...)))))))(((.((...(((......)))..(((((((((...(((........)))....))))))))).))..))).---------- ( -20.40) >DroSec_CAF1 11185 120 + 1 UUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUUAUUACUAUUUAUUGGCUUAUGAGUCUGGCAGAAAUGAUAU ...............(((((.((((((((...)))))))).((((.(((((......))...(((((((((...(((........)))....))))))))))))...))))))))).... ( -25.00) >DroSim_CAF1 11159 120 + 1 UUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUUAUUACUAUUUAUUGGCUUAUGAGUCUGGCAGAAAUGAUAU ...............(((((.((((((((...)))))))).((((.(((((......))...(((((((((...(((........)))....))))))))))))...))))))))).... ( -25.00) >consensus UUUUUUCCACAUACACAUUUUCCGAUAAGUCCCUUAUCGGCCUGCAACUGAACUUCAUCACCCGUGGGCUAUUUAUAUUUAUUACUAUUUAUUGGCUUAUGAGUCUGGCAGAAAUGAUAU .....................((((((((...)))))))).((((.(((((......))...(((((((((...(((........)))....))))))))))))...))))......... (-21.33 = -21.67 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:47:24 2006