
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,818,482 – 15,818,575 |
| Length | 93 |
| Max. P | 0.732296 |

| Location | 15,818,482 – 15,818,575 |
|---|---|
| Length | 93 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 71.47 |
| Mean single sequence MFE | -27.43 |
| Consensus MFE | -3.23 |
| Energy contribution | -3.67 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.97 |
| Structure conservation index | 0.12 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.732296 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
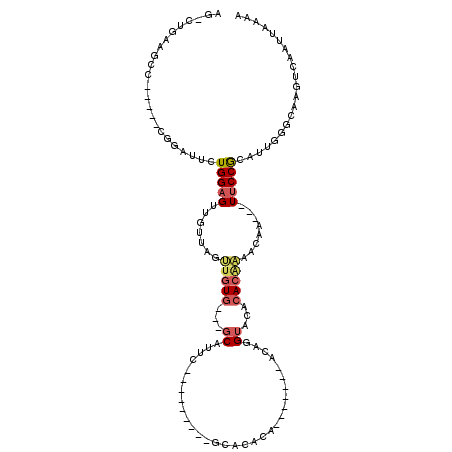
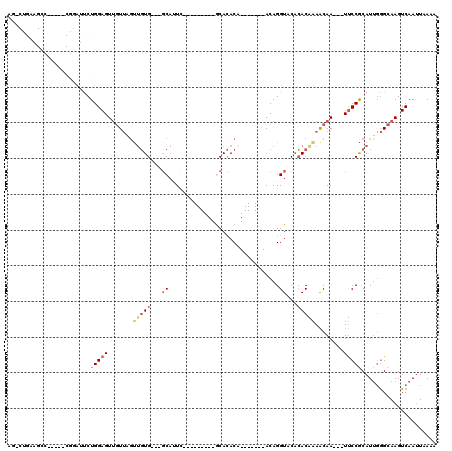
>2L_DroMel_CAF1 15818482 93 + 22407834 AGACUGAAGCC-----CGGAUUCUGGAGUUGUUAGUUGUG---GCAUUC---------GCACACA-------ACAGGUACACACAAAACAA---UUCCACAUUGGGCAAGUCAAUUAAAA .((((...(((-----(((....((((((((((.((((((---((....---------)).))))-------)).(.....)....)))))---)))))..)))))).))))........ ( -34.50) >DroPse_CAF1 11612 114 + 1 ---CGGAAGCCUUUGCCAAUGUCUGGAGUGGUGCGCUUUG---CCACUCCUCGGGUCUACAUACGGUUACACACAGGUACCAACGGAGCAAAAUUCCCGCAUGUAGCAAGUCAAUUAAAA ---.((((...(((((...(((..((((((((.......)---)))))))..............(((.((......))))).)))..))))).)))).((.....))............. ( -27.40) >DroSec_CAF1 13078 93 + 1 AGACUGAAGCC-----CGGAUUCUGGAGUUGUUAGUUGUG---GCAUUC---------GCACACA-------ACAGGUACACACAAAACAA---UUCCACAUUGGGCAAGUCAAUUAAAA .((((...(((-----(((....((((((((((.((((((---((....---------)).))))-------)).(.....)....)))))---)))))..)))))).))))........ ( -34.50) >DroSim_CAF1 13086 93 + 1 AGACUGAAGCC-----CGGAUUCUGGAGUUGUUAGUUGUG---GCAUUC---------ACACACA-------ACAGGUACACACAAAACAA---UUCCGCAUUGGGCAAGUCAAUUAAAA .((((...(((-----(((....((((((((((.((((((---......---------...))))-------)).(.....)....)))))---)))))..)))))).))))........ ( -31.00) >DroEre_CAF1 12839 79 + 1 AG-------------------UCUGGAGUUGUUCGUUGUG---GCAUUU---------GCACACA-------ACAGGUACACACAAAACAA---UUCCGCAUUGGGCAAGUCAAUUAAAA .(-------------------((((((((((((.((((((---((....---------)).))))-------)).(.....)....)))))---)))).....))))............. ( -21.90) >DroYak_CAF1 38592 82 + 1 AG-------------------CCUGGAGUUAUUAACAGUGUCUGCAAUU---------GCACACA-------ACAGGUACACACAAAACAA---UUCCGCAUUGGGCAAGUCAAAUCAAU .(-------------------(((((((((.......((((.(((..((---------(....))-------)...))).)))).....))---)))).....))))............. ( -15.30) >consensus AG_CUGAAGCC_____CGGAUUCUGGAGUUGUUAGUUGUG___GCAUUC_________GCACACA_______ACAGGUACACACAAAACAA___UUCCGCAUUGGGCAAGUCAAUUAAAA .......................(((((.......(((((...((...............................))...)))))........)))))..................... ( -3.23 = -3.67 + 0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:56 2006