
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,778,486 – 15,778,587 |
| Length | 101 |
| Max. P | 0.743418 |

| Location | 15,778,486 – 15,778,587 |
|---|---|
| Length | 101 |
| Sequences | 4 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 71.08 |
| Mean single sequence MFE | -21.85 |
| Consensus MFE | -11.66 |
| Energy contribution | -11.22 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.743418 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
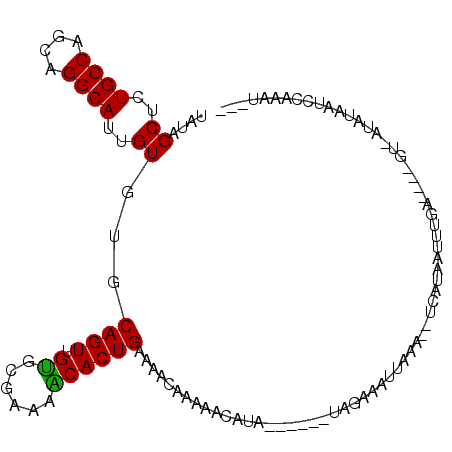
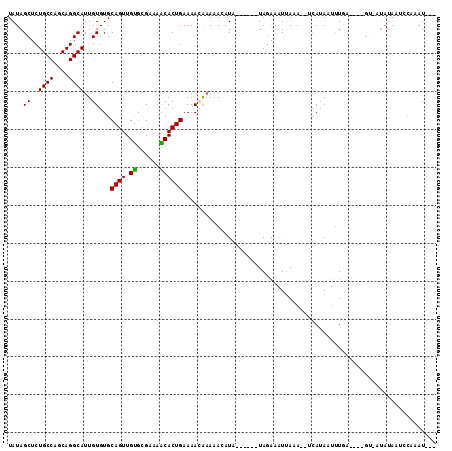
>2L_DroMel_CAF1 15778486 101 - 22407834 UAUAGCUCUGCCAGCAGGCAUUGUGUGCAGUUGCGCGAAAGCACUGAAAAGAAAAAGGUA------UAUGAAUUAAA--UCAUAAUUUGAUACAGU-AUAUAAACCACAU--- ....((.((((..((((...))))..))))..))((....)).((....)).....(((.------((((.(((..(--(((.....))))..)))-.)))).)))....--- ( -22.40) >DroSim_CAF1 29963 95 - 1 UAUAGCUCUGCCAGCAGGCAUUGUGUGCAGUUGUGCGAAAACACUGAAAAGAAAUAAGUA------UAGAAAUUAAA--UCAUAAUUUU------U-GUAAAAGGCUUAU--- ...((((.(((((((..((((...)))).)))).)))......((....)).......((------(((((((((..--...)))))))------)-)))...))))...--- ( -18.20) >DroEre_CAF1 31801 102 - 1 UAUAGCUCUGCCAGCAGGCAUUGUGUGCAGUUGU-UGAAAGCACUGAAAACGCAAACAUUUAAGAUUAGAAAUUAGA--UCACAUUUUGA----G----CUGUUCGAAAUACU .(((((((((((....))))((((((.((((.((-.....))))))...))))))........((((........))--)).......))----)----)))).......... ( -25.60) >DroYak_CAF1 31153 110 - 1 UAUAGCUCUGCCAGCAGGCAUUGUGUGCAGUUGUGCGAAAACACUGAAAACGAAAACAUUUAAAAUUAGAAAUAAAAAGUUAAAUGCUAA---AGCAAUAUAUUCCAAAUGGA ....(((.((((....))))((((...((((.((......))))))...))))...(((((((..(((....)))....)))))))....---))).......(((....))) ( -21.20) >consensus UAUAGCUCUGCCAGCAGGCAUUGUGUGCAGUUGUGCGAAAACACUGAAAACAAAAACAUA______UAGAAAUUAAA__UCAUAAUUUGA____GU_AUAUAAUCCAAAU___ ....((..((((....))))..))...((((.((......))))))................................................................... (-11.66 = -11.22 + -0.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:46:08 2006