
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,663,016 – 15,663,134 |
| Length | 118 |
| Max. P | 0.978214 |

| Location | 15,663,016 – 15,663,134 |
|---|---|
| Length | 118 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 62.40 |
| Mean single sequence MFE | -29.97 |
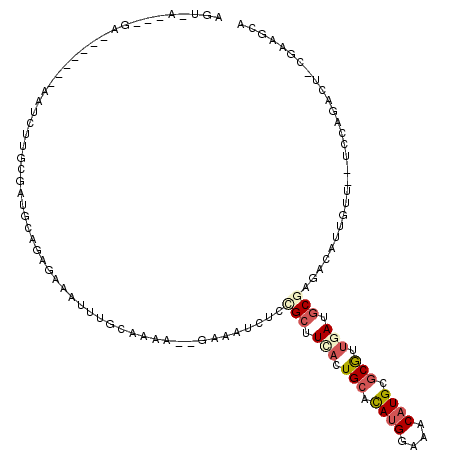
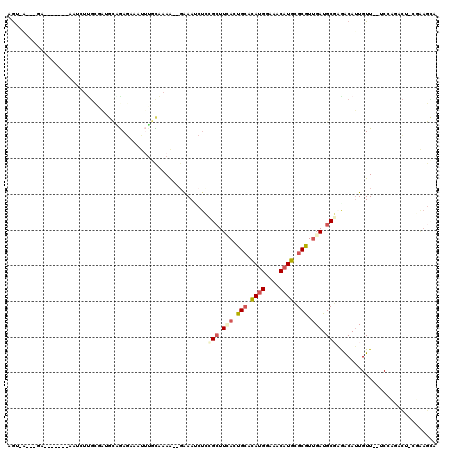
| Consensus MFE | -12.16 |
| Energy contribution | -13.05 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.31 |
| Mean z-score | -2.37 |
| Structure conservation index | 0.41 |
| SVM decision value | 1.81 |
| SVM RNA-class probability | 0.978214 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
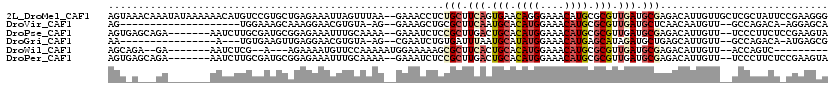
>2L_DroMel_CAF1 15663016 118 - 22407834 AGUAAACAAAUAUAAAAAACAUGUCCGUGCUGAGAAAUUAGUUUAA--GAAACCUCUGCUUCAGUGAACAGGGAAACAUGCGCGUUGAUGCGAGACAUUGUUGCUCGCUAUUCCGAAGGG ....................((((.((((((((....)))).....--.....(((((.(((...))))))))...)))).))))....(((((.((....)))))))....((....)) ( -23.80) >DroVir_CAF1 43087 94 - 1 AG--------------------UGGAAAGCAAAGGAACGUGUA-AG--GAAAGCUGCGCUUCAAUGCACAUGGAAACAUGCGCGUUGAUGCUCAACAAUGUU--GCCAGACA-AGGAGCA ..--------------------((...((((.......(((((-..--......))))).(((((((.((((....)))).)))))))))))...)).((((--.((.....-.)))))) ( -28.80) >DroPse_CAF1 22379 109 - 1 AGUGAGCAGA-------AAUCUUGCGAUGCGGAGAAAUUUGCAAAA--GAAAUCUCCGCUUGACUGCACAUGGAAACAUGCGCGUUGAUGCGAGACAUUGUU--UCCCUUCUCCGAAGUA .(..(((((.-------..(((((((..(((((((..(((.....)--))..)))))))(..((.((.((((....)))).))))..))))))))..)))))--..)((((...)))).. ( -37.50) >DroGri_CAF1 25516 95 - 1 AA----------------A---UGUGAAGUUGAGGAACGUGUA-AG--CGAAUCUGUGAUUUAAUGCAUAUGGAAACAUGAGCAUAGAUGCUGAGCAUUGUU--GCCAGACA-AUGAGCG ..----------------.---.((...((((.....(((...-.)--))..((((.(...((((((.((((....))))((((....))))..))))))..--.)))))))-))..)). ( -24.60) >DroWil_CAF1 43028 95 - 1 AGCAGA--GA-------AAUCUCG--A---AGAAAAUGUUCCAAAAAUGGAAAAAGCGCUUCACUGCACAUGGAAACAUGCGCGUUGAUGCGAGACAUUGUU--ACCAGUC--------- (((.((--(.-------...))).--.---....(((((((((....)))).....(((.(((.(((.((((....)))).))).))).)))..))))))))--.......--------- ( -27.60) >DroPer_CAF1 20079 109 - 1 AGUGAGCAGA-------AAUCUUGCGAUGCGGAGAAAUUUGCAAAA--GAAAUCUCCGCUUGACUGCACAUGGAAACAUGCGCGUUGAUGCGAGACAUUGUU--UCCCUUCUCCGAAGUA .(..(((((.-------..(((((((..(((((((..(((.....)--))..)))))))(..((.((.((((....)))).))))..))))))))..)))))--..)((((...)))).. ( -37.50) >consensus AGU_A___GA_______AAUCUUGCGAUGCAGAGAAAUUUGCAAAA__GAAAUCUCCGCUUCACUGCACAUGGAAACAUGCGCGUUGAUGCGAGACAUUGUU__UCCAGACU_CGAAGCA ........................................................(((.(((.(((.((((....)))).))).))).)))............................ (-12.16 = -13.05 + 0.89)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:45:17 2006