
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,599,058 – 15,599,149 |
| Length | 91 |
| Max. P | 0.624982 |

| Location | 15,599,058 – 15,599,149 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 79.79 |
| Mean single sequence MFE | -14.17 |
| Consensus MFE | -7.76 |
| Energy contribution | -8.38 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.19 |
| SVM RNA-class probability | 0.624982 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
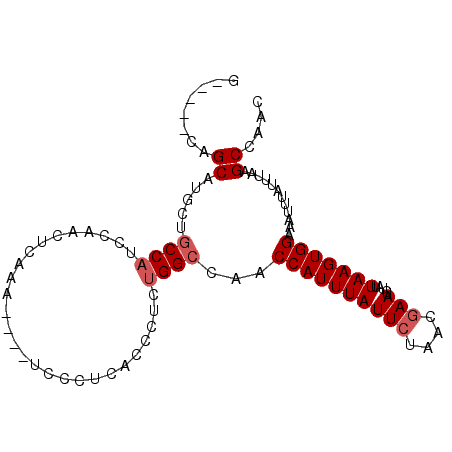
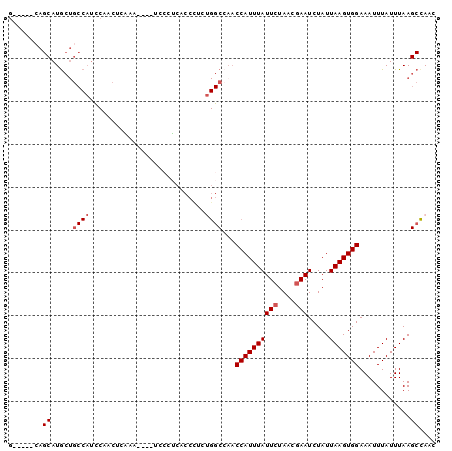
>2L_DroMel_CAF1 15599058 91 - 22407834 G-----CAGCAUGCUGCCAUCCAACUCAAA----UCCCUCACCCUCAGGCCAACCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCCAAC (-----(((....)))).............----.............(((...((((((((((....))).....)))))))(((....)))..)))... ( -14.60) >DroSec_CAF1 105695 91 - 1 G-----AAGCAUGCUGCCAUCCAACUCAAA----UCCCUCACCUUCUGGCAAACCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCCAAC .-----..((....(((((...........----............)))))..((((((((((....))).....)))))))............)).... ( -13.20) >DroSim_CAF1 97989 91 - 1 G-----CAGCAUGCUGCCAUCCAACUCGAA----UCCCUCACCCUCUGGCCAACCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCCAAC (-----(((....)))).............----............((((...((((((((((....))).....)))))))(((....)))..)))).. ( -15.20) >DroEre_CAF1 99543 82 - 1 G-----CAGCAUGCUGCCAUCCCACU-------------CACCCUUUGGCCAACCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCCAAC (-----(((....)))).........-------------......(((((...((((((((((....))).....)))))))(((....)))..))))). ( -15.90) >DroYak_CAF1 103129 95 - 1 G-----CAGCAUGCUGCCAUUCCACUUACCCUCUCACCUCACCCUCUGGCCUUCCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCUAAC (-----(((....))))..(((((((((..................(((....)))...((((....))))....)))))))))................ ( -19.10) >DroAna_CAF1 320973 81 - 1 ACAACCGAGCAUA-----------CUCUAA----GAACGCCCCCUU---CCAACCAUUUAUUGUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGC-AAA ......(((....-----------)))...----............---....(((((((..(........)...)))))))..............-... ( -7.00) >consensus G_____CAGCAUGCUGCCAUCCAACUCAAA____UCCCUCACCCUCUGGCCAACCAUUUAUUCUAACGAAUCUAUUAAGUGGAAAUUUAUUUAAGCCAAC ........((.....((((...........................))))...((((((((((....))).....)))))))............)).... ( -7.76 = -8.38 + 0.62)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:54 2006