
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,581,927 – 15,582,018 |
| Length | 91 |
| Max. P | 0.500000 |

| Location | 15,581,927 – 15,582,018 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 75.18 |
| Mean single sequence MFE | -24.00 |
| Consensus MFE | -12.91 |
| Energy contribution | -12.80 |
| Covariance contribution | -0.11 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.54 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15581927 91 - 22407834 AGC-GAGGAAGCUAG--GAAACUGA--AGCUGAAA----------------CAGGUAGCAGCUGCC---UGGCGCUAA-AGUGCCAACUUAUGCCA--CGAAAAUGCUUGACUAUAAU (((-((((.((((((--....))..--.((((...----------------....)))))))).))---)(((((...-.)))))...........--......)))).......... ( -24.40) >DroVir_CAF1 115419 105 - 1 -CAGGCGAGAGCU----GAAACUGU--AACUGAAACA----GCAACUG-AACAGGUAGCAGCUGCCUCCUGACGCUAA-AGUGCCAACUUAUGCCACACAAAAAUGCUUGAAUAUAAU -((((((..(((.----....((((--.......)))----)......-..((((..((....))..))))..)))..-.(((.((.....)).))).......))))))........ ( -23.10) >DroPse_CAF1 142938 107 - 1 AGA-GGGAGAGCUGG--GAAACUAAAAAACUGAAACAGGUUGCA--UGGAGCAGGCAGCAGGUGCC---UGGCGCUAA-AGUGCCAACUUAUGCCA--CAAAAAUGCUUGAAUAUAAU ...-....(((((((--....)))..............((.(((--(((...(((((.....))))---)(((((...-.)))))...)))))).)--)......))))......... ( -28.30) >DroYak_CAF1 93250 91 - 1 CAC-GAUGGAGCUGG--CAAGCUGA--AACUGAAA----------------CAGGUAGCAGCUGCC---UGGCGCUAA-AGUGCCAACUUAUGCCA--CGAAAAUGCUUGACUAUAAU ..(-(.(((....((--((.((((.--.(((....----------------..)))..))))))))---((((((...-.)))))).......)))--)).................. ( -28.60) >DroMoj_CAF1 123885 95 - 1 -GAGGCCAAAGCU----GAAACUGU--AACUGGA---------------AACAGGUAGCAGCUGCCUCCUGACACUAA-AGUGCCAACUUAUGCCACACAAAAAUGCUUGAAUAUAAU -(((((...((((----(...(((.--..(((..---------------..))).)))))))))))))..........-.(((.((.....)).)))..................... ( -22.20) >DroAna_CAF1 240897 94 - 1 ACC-AGGAGAGGAGGUAAAAACUGA--AAUUGAAA----------------CAGGUAGCAGCUGCC---UGGUGCUAAAAGUGCUAACUUAUUCCA--CGAAAAUGCUUGAAUAUAAU ...-.((((...((((.........--........----------------(((((((...)))))---))(..(.....)..)..)))).)))).--.................... ( -17.40) >consensus AGA_GAGAGAGCUGG__GAAACUGA__AACUGAAA________________CAGGUAGCAGCUGCC___UGGCGCUAA_AGUGCCAACUUAUGCCA__CAAAAAUGCUUGAAUAUAAU ........((((.........................................(((((...)))))...((((((.....))))))...................))))......... (-12.91 = -12.80 + -0.11)

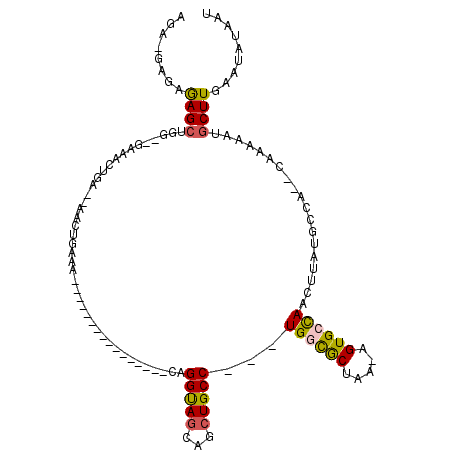

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:50 2006