
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,541,622 – 15,541,720 |
| Length | 98 |
| Max. P | 0.814898 |

| Location | 15,541,622 – 15,541,720 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 90.17 |
| Mean single sequence MFE | -23.47 |
| Consensus MFE | -19.78 |
| Energy contribution | -20.67 |
| Covariance contribution | 0.89 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.793717 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15541622 98 + 22407834 GUGAGGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU-UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.((((((((((-((.((......)).)))..))))))))))))))......))..)).))..... ( -25.32) >DroSec_CAF1 48527 98 + 1 GCGAAGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU-UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.((((((((((-((.((......)).)))..))))))))))))))......))..)).))..... ( -25.32) >DroSim_CAF1 47171 98 + 1 GCGAAGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU-UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.((((((((((-((.((......)).)))..))))))))))))))......))..)).))..... ( -25.32) >DroEre_CAF1 49180 98 + 1 GUGAAGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU-UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.((((((((((-((.((......)).)))..))))))))))))))......))..)).))..... ( -25.32) >DroYak_CAF1 51320 98 + 1 GUGAAGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU-UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.((((((((((-((.((......)).)))..))))))))))))))......))..)).))..... ( -25.32) >DroAna_CAF1 163460 85 + 1 GGGA-------UGUGC---GGCAAACAAAUCCUGUAGCAUAAUUUUUUGGCAAGAACGUAAAAUUAAUUAUGCGAGAAAAGAAGAGCGGGAGAUG---- ....-------.....---......((..((((((.((((((((.(((.((......)).)))..))))))))............))))))..))---- ( -14.22) >consensus GUGAAGGGAAAUGAGCGUGGGCAAACAAAUCCUGUCGCAUAAUUU_UUGGCAAGAACGUGAAAUUAAUUAUGCGCAGGAAGAAAAGCGGCAGGCGAAUG ..............((.((.((.......(((((.(((((((((..((.((......)).))...))))))))))))))......))..)).))..... (-19.78 = -20.67 + 0.89)
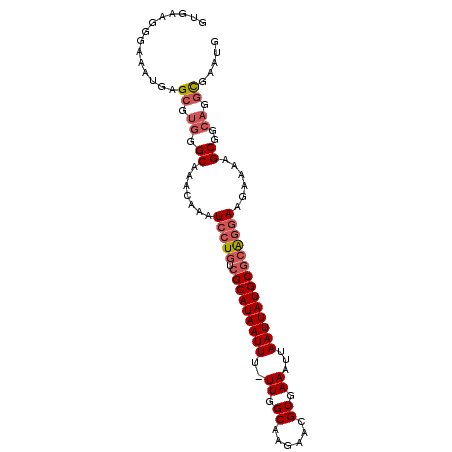



| Location | 15,541,622 – 15,541,720 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 90.17 |
| Mean single sequence MFE | -19.03 |
| Consensus MFE | -15.37 |
| Energy contribution | -15.95 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.15 |
| Mean z-score | -2.60 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814898 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15541622 98 - 22407834 CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA-AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCCUCAC .....((.((..((......((((((((((((((.....((......))....-.))))))))).))))).......)).)).)).............. ( -20.62) >DroSec_CAF1 48527 98 - 1 CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA-AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCUUCGC .....((.((..((......((((((((((((((.....((......))....-.))))))))).))))).......)).)).)).............. ( -20.62) >DroSim_CAF1 47171 98 - 1 CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA-AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCUUCGC .....((.((..((......((((((((((((((.....((......))....-.))))))))).))))).......)).)).)).............. ( -20.62) >DroEre_CAF1 49180 98 - 1 CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA-AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCUUCAC .....((.((..((......((((((((((((((.....((......))....-.))))))))).))))).......)).)).)).............. ( -20.62) >DroYak_CAF1 51320 98 - 1 CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA-AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCUUCAC .....((.((..((......((((((((((((((.....((......))....-.))))))))).))))).......)).)).)).............. ( -20.62) >DroAna_CAF1 163460 85 - 1 ----CAUCUCCCGCUCUUCUUUUCUCGCAUAAUUAAUUUUACGUUCUUGCCAAAAAAUUAUGCUACAGGAUUUGUUUGCC---GCACA-------UCCC ----......................((((((((..((((..((....)).))))))))))))....((((.(((.....---))).)-------))). ( -11.10) >consensus CAUUCGCCUGCCGCUUUUCUUCCUGCGCAUAAUUAAUUUCACGUUCUUGCCAA_AAAUUAUGCGACAGGAUUUGUUUGCCCACGCUCAUUUCCCUUCAC .....((.((..((......((((((((((((((.....................))))))))).))))).......)).)).)).............. (-15.37 = -15.95 + 0.59)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:44:36 2006