
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,677,855 – 1,677,968 |
| Length | 113 |
| Max. P | 0.919533 |

| Location | 1,677,855 – 1,677,968 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 84.69 |
| Mean single sequence MFE | -33.77 |
| Consensus MFE | -24.17 |
| Energy contribution | -24.05 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.709359 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
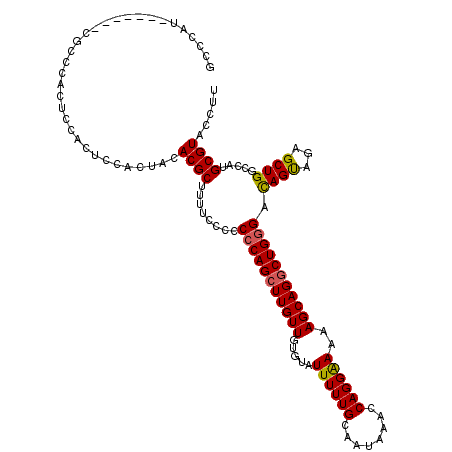
>2L_DroMel_CAF1 1677855 113 + 22407834 GCCCAU-------CGCCCACUCCAAUCCACUAUACGCUUUUCCCCCCCAGCUUGUUGUGUAUUUUUGCAAUAAACCAGGAAAAAGCAGACUGGGACAGUAGAGCUGGCCAUGCGUACCUU ((.(((-------.(((..(((.............((((((.((.....(.((((((((......)))))))).)..)).))))))..((((...)))).)))..))).))).))..... ( -23.80) >DroSec_CAF1 4223 104 + 1 C--------------UCCACUCCACUCUACUACACGC--UUUUCCCCCAGCUUGUUGUGUAUUUUUGCAAUAAACCAGGAAAAAGCAGGCUGGGACAGUGGAGCUGGCCAUGCGUACCUU .--------------.((((((((((...(.....).--......(((((((((((.....((((((........))))))..)))))))))))..))))))).)))............. ( -30.50) >DroSim_CAF1 5214 120 + 1 GCCCAUCGCCCAUCGCCCACUCCACUCCACUACACGCUCUUUUCCCCCAGCUUGUUGUGUAUUUUUGCAAUAAACCAGGGAAAAGCAGGCUGGGACAGUGGAGCUGGCCAUGCGUACCUU .......((.(((.(((..(((((((...(((...(((((((((((...(.((((((((......)))))))).)..))))))))..))))))...)))))))..))).))).))..... ( -38.90) >DroYak_CAF1 5294 113 + 1 GCCCAU-------GGCCCACUCCACCUCACUGCACGCUUUUCCCCCGCAGCUUGUUGUGUAUUUUUGCAAUAAAGCAGGGAAAAGCAGGCUGGGAUAGCAGAGCUGGCCAUGCGUACCUU ((.(((-------((((..(((....((...((..(((((((((..((...((((((((......)))))))).)).)))))))))..))...)).....)))..))))))).))..... ( -41.90) >consensus GCCCAU_______CGCCCACUCCACUCCACUACACGCUUUUCCCCCCCAGCUUGUUGUGUAUUUUUGCAAUAAACCAGGAAAAAGCAGGCUGGGACAGUAGAGCUGGCCAUGCGUACCUU .................................((((........(((((((((((.....((((((........))))))..))))))))))).((((...)))).....))))..... (-24.17 = -24.05 + -0.12)


| Location | 1,677,855 – 1,677,968 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 84.69 |
| Mean single sequence MFE | -41.90 |
| Consensus MFE | -29.62 |
| Energy contribution | -29.00 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.919533 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
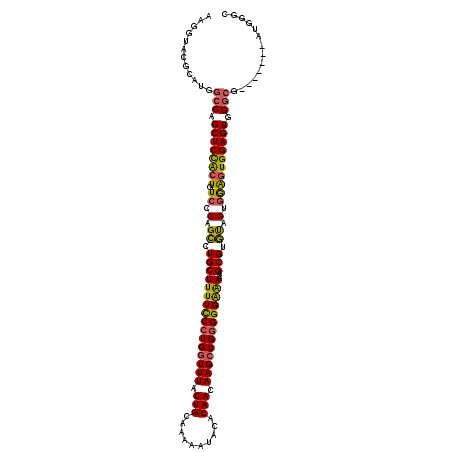
>2L_DroMel_CAF1 1677855 113 - 22407834 AAGGUACGCAUGGCCAGCUCUACUGUCCCAGUCUGCUUUUUCCUGGUUUAUUGCAAAAAUACACAACAAGCUGGGGGGGAAAAGCGUAUAGUGGAUUGGAGUGGGCG-------AUGGGC ......(.(((.(((.((((((..(((((.((.((((((((((((((((.(((..........))).)))))...))))))))))).)).).)))))))))).))).-------))).). ( -39.50) >DroSec_CAF1 4223 104 - 1 AAGGUACGCAUGGCCAGCUCCACUGUCCCAGCCUGCUUUUUCCUGGUUUAUUGCAAAAAUACACAACAAGCUGGGGGAAAA--GCGUGUAGUAGAGUGGAGUGGA--------------G ..(((.......))).((((((((.(.(..((.((((((((((..((((.(((..........))).))))..).))))))--))).)).).).))))))))...--------------. ( -33.40) >DroSim_CAF1 5214 120 - 1 AAGGUACGCAUGGCCAGCUCCACUGUCCCAGCCUGCUUUUCCCUGGUUUAUUGCAAAAAUACACAACAAGCUGGGGGAAAAGAGCGUGUAGUGGAGUGGAGUGGGCGAUGGGCGAUGGGC ......(.(((.(((.(((((((..((((.((.((((((((((..((((.(((..........))).))))..)))))...))))).)).).)))))))))).))).))).)........ ( -46.70) >DroYak_CAF1 5294 113 - 1 AAGGUACGCAUGGCCAGCUCUGCUAUCCCAGCCUGCUUUUCCCUGCUUUAUUGCAAAAAUACACAACAAGCUGCGGGGGAAAAGCGUGCAGUGAGGUGGAGUGGGCC-------AUGGGC ......(.(((((((.((((..((.((.(.((.(((((((((((((......)).......(.((......)).)))))))))))).)).).))))..)))).))))-------))).). ( -48.00) >consensus AAGGUACGCAUGGCCAGCUCCACUGUCCCAGCCUGCUUUUCCCUGGUUUAUUGCAAAAAUACACAACAAGCUGGGGGAAAAAAGCGUGUAGUGGAGUGGAGUGGGCG_______AUGGGC ............(((.((((((((.((.(.((.((((((((((((((((.(((..........))).)))))))))))))...))).)).).)))))))))).))).............. (-29.62 = -29.00 + -0.62)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:54 2006