
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,454,626 – 15,454,717 |
| Length | 91 |
| Max. P | 0.955063 |

| Location | 15,454,626 – 15,454,717 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -43.66 |
| Consensus MFE | -28.66 |
| Energy contribution | -29.46 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.66 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15454626 91 + 22407834 UUGGCCACACAGGGCGAGCAGGGGCCAGUUGAGCGGUGCCCAGU--------UGCCCAGUUGGCCACUUGGCAACUCGGCCUG-CCGAGAGAUGUGUGCU ..(((((((((((.(((((((((((((..((.((..(.....).--------.)).))..))))).))))....)))).))))-.(....).)))).))) ( -39.10) >DroSec_CAF1 192678 81 + 1 UUGGCCACACAGGGCGAGCAGGGGCCAGUUGAGCGGUGCCC----------------AGUUGGCCACUUGGCAACUCGUCCUG-CCGA--GAUGUGUGCU ..(((((((((((((((((((((((((..((.((...)).)----------------)..))))).))))....)))))))))-....--..)))).))) ( -37.00) >DroSim_CAF1 198610 89 + 1 UUGGCCACACAGGGCGUGCAGGGGCCAGUUGAGCGGUGCCCAGU--------UGCCCAGUUGGCCACUUGGCAACUCGGCCUG-CCGA--GAUGUGUGCU .((((((.((.(((((.((.((((((........))).))).))--------))))).))))))))....(((.(((((....-))))--).)))..... ( -39.20) >DroEre_CAF1 202936 97 + 1 UUGGCCACACAGGGCGAGCAGGGGCCAGUUGAGCGGUGCCCAGUUGCCCAGUUGCCCAGUUGGCCACUUGGCAACUGGGCCUG-CCGA--GAUGUGUGGU ...((((((((.(((........)))..(((.((((.(((((((((((.(((.(((.....))).))).))))))))))))))-))))--..)))))))) ( -56.00) >DroYak_CAF1 200222 98 + 1 UUGGCCACACAGGGCGAGCAGGGGCCAGUUGAGCGGUACCCAGUUGCCGAGUUGCCCAGUUGGCCACUUGGCAACUCGGCCUGGCCGA--GAUGUGUGCU ..(((.(((((.(((.((..((((((........))).)))....(((((((((((.(((.....))).))))))))))))).)))..--..)))))))) ( -47.00) >consensus UUGGCCACACAGGGCGAGCAGGGGCCAGUUGAGCGGUGCCCAGU________UGCCCAGUUGGCCACUUGGCAACUCGGCCUG_CCGA__GAUGUGUGCU .....((((((...((.((((((((..(.....)...))))............(((.(((((.((....))))))).)))))).))).....)))))).. (-28.66 = -29.46 + 0.80)



| Location | 15,454,626 – 15,454,717 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 88.20 |
| Mean single sequence MFE | -41.88 |
| Consensus MFE | -29.50 |
| Energy contribution | -30.14 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -3.12 |
| Structure conservation index | 0.70 |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.955063 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
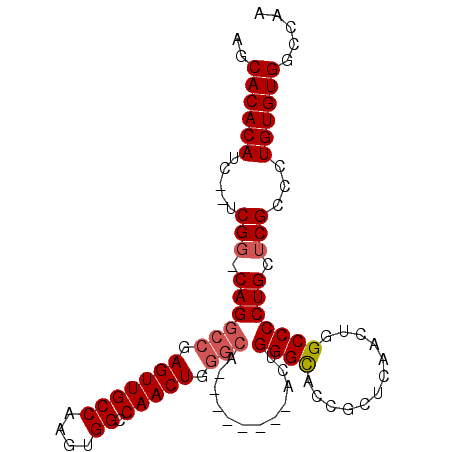
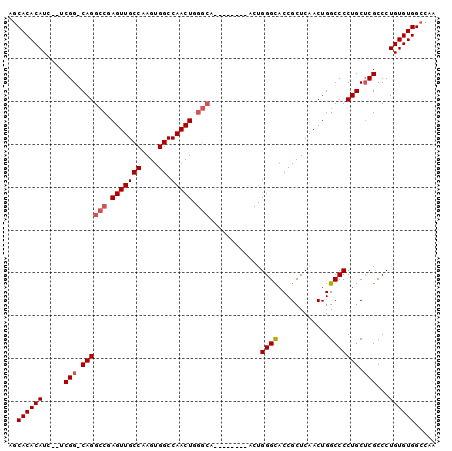
>2L_DroMel_CAF1 15454626 91 - 22407834 AGCACACAUCUCUCGG-CAGGCCGAGUUGCCAAGUGGCCAACUGGGCA--------ACUGGGCACCGCUCAACUGGCCCCUGCUCGCCCUGUGUGGCCAA .((.(((((....(((-((((((.(((((((.(((.....))).))))--------))).)))...(((.....)))..)))).))....)))))))... ( -38.60) >DroSec_CAF1 192678 81 - 1 AGCACACAUC--UCGG-CAGGACGAGUUGCCAAGUGGCCAACU----------------GGGCACCGCUCAACUGGCCCCUGCUCGCCCUGUGUGGCCAA ..((((((..--.(((-((((.(.(((((....(((.((....----------------)).)))....))))).)..))))).))...))))))..... ( -32.50) >DroSim_CAF1 198610 89 - 1 AGCACACAUC--UCGG-CAGGCCGAGUUGCCAAGUGGCCAACUGGGCA--------ACUGGGCACCGCUCAACUGGCCCCUGCACGCCCUGUGUGGCCAA ..((((((..--.(((-((((((.(((((((.(((.....))).))))--------))).)))...(((.....)))..)))).))...))))))..... ( -39.40) >DroEre_CAF1 202936 97 - 1 ACCACACAUC--UCGG-CAGGCCCAGUUGCCAAGUGGCCAACUGGGCAACUGGGCAACUGGGCACCGCUCAACUGGCCCCUGCUCGCCCUGUGUGGCCAA .(((((((..--.(((-((((((((((((((.(((.(((.....))).))).)))))))))))...(((.....)))..)))).))...))))))).... ( -52.10) >DroYak_CAF1 200222 98 - 1 AGCACACAUC--UCGGCCAGGCCGAGUUGCCAAGUGGCCAACUGGGCAACUCGGCAACUGGGUACCGCUCAACUGGCCCCUGCUCGCCCUGUGUGGCCAA ..((((((..--..(((((((((((((((((.(((.....))).)))))))))))...((((.....)))).))))))...(....)..))))))..... ( -46.80) >consensus AGCACACAUC__UCGG_CAGGCCGAGUUGCCAAGUGGCCAACUGGGCA________ACUGGGCACCGCUCAACUGGCCCCUGCUCGCCCUGUGUGGCCAA ..((((((.....(((.((((((.(((((((....)).))))).)))............((((............))))))).)))...))))))..... (-29.50 = -30.14 + 0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:44 2006