
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,413,433 – 15,413,526 |
| Length | 93 |
| Max. P | 0.999328 |

| Location | 15,413,433 – 15,413,526 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 96.77 |
| Mean single sequence MFE | -23.62 |
| Consensus MFE | -22.84 |
| Energy contribution | -23.44 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.585761 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15413433 93 + 22407834 AAUGCGGGCAACCUGAAGAUACCAAAAAAGCCAAAUAAAAGCGACGGAUGCGUGUCGCUUUCGACUGGCUCGGCCACUGAUUUAAUUCGGUAA ...((((....))...............(((((....(((((((((......)))))))))....)))))..)).(((((......))))).. ( -27.40) >DroSec_CAF1 159242 92 + 1 AAUGCGGGCAACCUGAAGAUACCAAA-AAGCCAAAUAAAAGCGACGGAUGCGUGUCGCUUUCGACUGGCUCUGCCACUGAUUUAAUUCGGUAA ...((((....)).............-.(((((....(((((((((......)))))))))....)))))..)).(((((......))))).. ( -27.40) >DroSim_CAF1 164233 93 + 1 AAUGCGGGCAACCUGAAGAUACCAAAAAAGCCAAAUAAAAGCGACGGAUGCGUGUCGCUUUCGACUGGCUCUGCCACUGAUUUAAUUCGGUAA ...((((....))...............(((((....(((((((((......)))))))))....)))))..)).(((((......))))).. ( -27.40) >DroEre_CAF1 165967 93 + 1 AAUGCGGGCAACCUGAAGAUACCAAAAAAGCCAACUAAAAGCGACGGAUGCGUGUCUCUUUCGACUGACUCUGCCACUGAUUUAAUUCGGUUG .....((....))...............((((.......((.(.((((..((.(((......)))))..)))).).))((......)))))). ( -15.60) >DroYak_CAF1 166380 93 + 1 AAUGCGGGCAACCUGAAGAUACCAAAAAAGCCAAAUAAAAGCGACGGAUGCGUGUCGCUUUCGACUGACUCUGCCACUGAUUUAAUUCGGUAA ...((((....))...............((.((....(((((((((......)))))))))....)).))..)).(((((......))))).. ( -20.30) >consensus AAUGCGGGCAACCUGAAGAUACCAAAAAAGCCAAAUAAAAGCGACGGAUGCGUGUCGCUUUCGACUGGCUCUGCCACUGAUUUAAUUCGGUAA ...((((....))...............(((((....(((((((((......)))))))))....)))))..)).(((((......))))).. (-22.84 = -23.44 + 0.60)

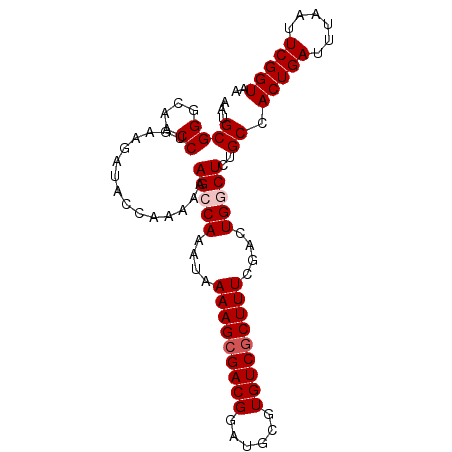
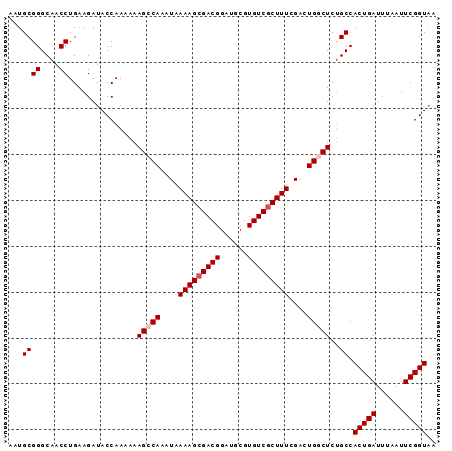
| Location | 15,413,433 – 15,413,526 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 96.77 |
| Mean single sequence MFE | -28.20 |
| Consensus MFE | -26.90 |
| Energy contribution | -27.06 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.95 |
| SVM decision value | 3.51 |
| SVM RNA-class probability | 0.999328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15413433 93 - 22407834 UUACCGAAUUAAAUCAGUGGCCGAGCCAGUCGAAAGCGACACGCAUCCGUCGCUUUUAUUUGGCUUUUUUGGUAUCUUCAGGUUGCCCGCAUU ................(..((((((((((..(((((((((........)))))))))..)))))))...(((.....))))))..)....... ( -30.10) >DroSec_CAF1 159242 92 - 1 UUACCGAAUUAAAUCAGUGGCAGAGCCAGUCGAAAGCGACACGCAUCCGUCGCUUUUAUUUGGCUU-UUUGGUAUCUUCAGGUUGCCCGCAUU ................(((((((((((((..(((((((((........)))))))))..)))))))-(((((.....))))).)).))))... ( -29.70) >DroSim_CAF1 164233 93 - 1 UUACCGAAUUAAAUCAGUGGCAGAGCCAGUCGAAAGCGACACGCAUCCGUCGCUUUUAUUUGGCUUUUUUGGUAUCUUCAGGUUGCCCGCAUU ................(((((((((((((..(((((((((........)))))))))..))))))).(((((.....))))).)).))))... ( -29.70) >DroEre_CAF1 165967 93 - 1 CAACCGAAUUAAAUCAGUGGCAGAGUCAGUCGAAAGAGACACGCAUCCGUCGCUUUUAGUUGGCUUUUUUGGUAUCUUCAGGUUGCCCGCAUU ((((((((....(((((....((((((((..(((((.(((........))).)))))..)))))))).)))))...))).)))))........ ( -24.50) >DroYak_CAF1 166380 93 - 1 UUACCGAAUUAAAUCAGUGGCAGAGUCAGUCGAAAGCGACACGCAUCCGUCGCUUUUAUUUGGCUUUUUUGGUAUCUUCAGGUUGCCCGCAUU ................(((((((((((((..(((((((((........)))))))))..))))))).(((((.....))))).)).))))... ( -27.00) >consensus UUACCGAAUUAAAUCAGUGGCAGAGCCAGUCGAAAGCGACACGCAUCCGUCGCUUUUAUUUGGCUUUUUUGGUAUCUUCAGGUUGCCCGCAUU ................(((((((((((((..(((((((((........)))))))))..))))))).(((((.....))))).)).))))... (-26.90 = -27.06 + 0.16)

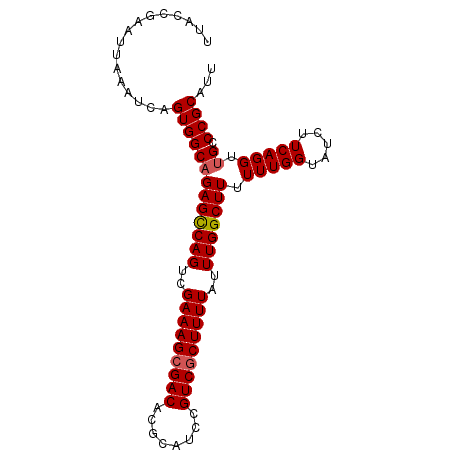
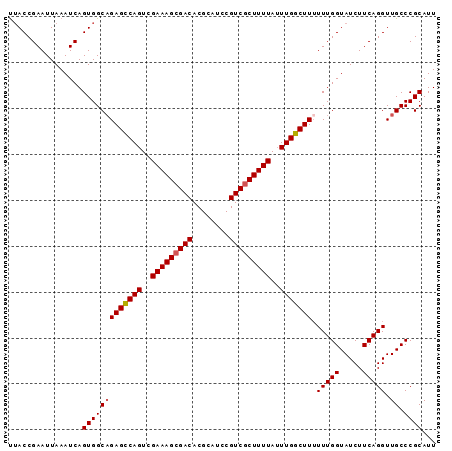
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:43:22 2006