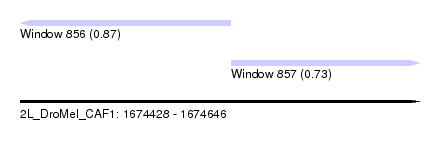
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,674,428 – 1,674,646 |
| Length | 218 |
| Max. P | 0.870535 |

| Location | 1,674,428 – 1,674,543 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.60 |
| Mean single sequence MFE | -26.63 |
| Consensus MFE | -24.55 |
| Energy contribution | -25.05 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.02 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1674428 115 - 22407834 AAAAACCAAUCAAGAGCAUCAAUAAAAGGCACGUCAAUCAGCGUGUGGCA-----AGGUAGGCUCCAGCCAUUUUCCUCCUCCUCGCUAAAACUGCUGCUCAACAAGCACGAAAAAAUCA .............(((((.((.......((((((......))))))(((.-----(((..(((....)))...........))).))).....)).)))))................... ( -23.72) >DroSec_CAF1 897 115 - 1 AAAAACCAAUCAAGAGCAUCAAUAAAAGGCACGUCAAUCAGCGUGUGGCA-----AGGUAGGCUCCAGCCAUUUUCCUCCUCCUCGGAAAAACUGCUGCUCAACAAGCACGAAAAAAUCA .............((((...........((((((......))))))((((-----.....(((....))).((((((........))))))..))))))))................... ( -27.30) >DroSim_CAF1 1785 115 - 1 AAAAACCAAUCAAGAGCAUCAAUAAAAGGCACGUCAAUCAGCGUGUGGCA-----AGGUAGGCUCCAGCCAUUUUCCUCCUCCUCGGAAAAACUGCUGCUCAACAAGCACGAAAAAAUCA .............((((...........((((((......))))))((((-----.....(((....))).((((((........))))))..))))))))................... ( -27.30) >DroYak_CAF1 1894 120 - 1 AAAAACCAAUCAAGAGCAUCAAUAAAAGGCACGUCAAUCAGCGUGUGGCAAGGCAAGGUAGGCUCCAGCCAUUUUCCUCCUGCUCGGAAAAACUGCUGCUCAACAAGCACGAAAAAAUCA .............((((...........((((((......))))))((((.(((..((......)).))).((((((........))))))..))))))))................... ( -28.20) >consensus AAAAACCAAUCAAGAGCAUCAAUAAAAGGCACGUCAAUCAGCGUGUGGCA_____AGGUAGGCUCCAGCCAUUUUCCUCCUCCUCGGAAAAACUGCUGCUCAACAAGCACGAAAAAAUCA .............((((...........((((((......))))))((((..........(((....))).((((((........))))))..))))))))................... (-24.55 = -25.05 + 0.50)

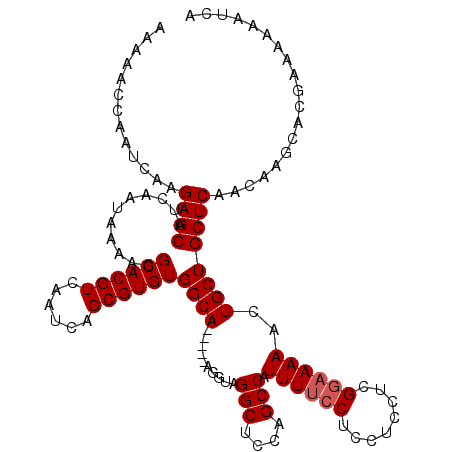
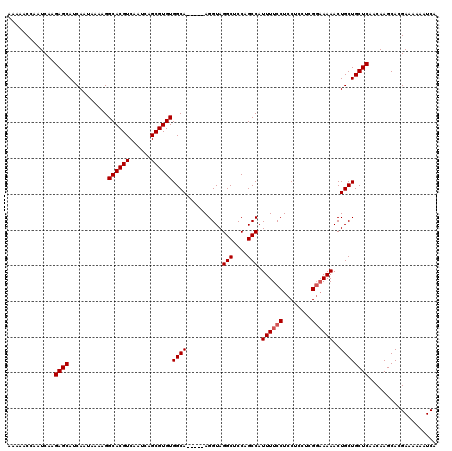
| Location | 1,674,543 – 1,674,646 |
|---|---|
| Length | 103 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.32 |
| Mean single sequence MFE | -21.51 |
| Consensus MFE | -18.05 |
| Energy contribution | -18.18 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.50 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.733072 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 1674543 103 + 22407834 ACCUUUUUACACAGACAUGCGGGCGGUGGCGCCACCUCUUUUCCAUCCACCUG-CCCACCUUUUUUUUUGUUCUCCUUC----------------ACCGUCCACCUGGUCGUUAGUUCAA .............(((....(((((((((.................))))).)-)))......................----------------..((.((....)).))...)))... ( -19.13) >DroSec_CAF1 1012 113 + 1 ACCUUUUAACACAGACAUGCGGGCGGUGG--CCUCCCCUUUUCCACACACCC--CCCACCUUUU--UUUGUUUUCCUUUAGAG-CCCAAUCACCCGCCGCCCACCUGGUCGUUAGUUCAA .....(((((...(((..(.(((((((((--.....................--..((......--..))..........((.-.....))..))))))))).)...))))))))..... ( -22.50) >DroSim_CAF1 1900 112 + 1 ACCUUUUUACACAGACAUGCGGGCGGUGG--CCUCCCCUUUUCCACACACCC---CCACCUUUU--UUUGUUCUCCUUUAGCG-CCCAACCACCCGCCGCCCACCUGGUCGUUAGUUCAA .............(((..(.(((((((((--.....................---.........--.................-.........))))))))).)...))).......... ( -20.94) >DroYak_CAF1 2014 109 + 1 ACCUUUUUACACAGACAUGCGGGCGGUGG--CCUCCCCUUUUACUC-CACCCCCUCCACUGUU--------UUUCCUUUAGUAUCGCAACCACCCGCCGCCCACCUGGUCGUUAGUUCAA .............(((..(.(((((((((--..........((((.-................--------........))))..........))))))))).)...))).......... ( -23.46) >consensus ACCUUUUUACACAGACAUGCGGGCGGUGG__CCUCCCCUUUUCCACACACCC__CCCACCUUUU__UUUGUUCUCCUUUAG_G_CCCAACCACCCGCCGCCCACCUGGUCGUUAGUUCAA .............(((..(.(((((((((................................................................))))))))).)...))).......... (-18.05 = -18.18 + 0.13)

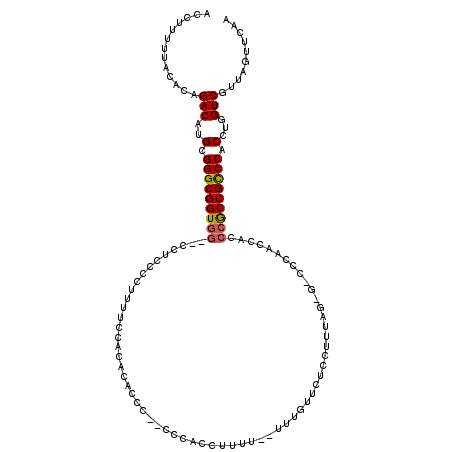
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:38:36 2006