
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 15,117,940 – 15,118,077 |
| Length | 137 |
| Max. P | 0.658983 |

| Location | 15,117,940 – 15,118,039 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 77.63 |
| Mean single sequence MFE | -14.66 |
| Consensus MFE | -11.86 |
| Energy contribution | -12.66 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.15 |
| Structure conservation index | 0.81 |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.658983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15117940 99 - 22407834 GCUAGCUAGCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAAAAAACCACCUACCCACCUACCCCGCAUCCACCACC (((((....)))))..(((((.((((.(....).)))).)))))....................................................... ( -16.70) >DroSec_CAF1 62732 82 - 1 GCUA----GCUAGCGAGCAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG----CAACCU--------ACCC-CCAUCCACCACC (((.----...)))..(((((.((((.(....).)))).)))))....................----......--------....-............ ( -11.70) >DroSim_CAF1 60419 82 - 1 GCUA----GCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG----CAACCU--------ACCC-CCAUCCACCACC (((.----...)))..(((((.((((.(....).)))).)))))....................----......--------....-............ ( -11.40) >DroEre_CAF1 63741 85 - 1 GCUGGCUAGCUAGCAAGCGGGCUAGACCUGUCGCUCUGUCCUGUAAA-CCAGCCAAAGGGAAAU-----CACCCA-------CCCA-CCAGCCACCACC (((((...(((.(...(((((.((((.(....).)))).)))))...-).)))....(((....-----..))).-------....-)))))....... ( -23.30) >DroYak_CAF1 77673 80 - 1 GCUAGCUAGCGAGCGA---GGCUAGACCUGUCGUUCUGUCCUGCAAAUCCACCCAAA---AAAC-----CACCUA-------CCUA-CCACCCACCACC (((....)))..((.(---((.((((.(....).)))).))))).............---....-----......-------....-............ ( -10.20) >consensus GCUAGCUAGCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG____CCACCUA_______ACCC_CCAUCCACCACC (((((....)))))..(((((.((((.(....).)))).)))))....................................................... (-11.86 = -12.66 + 0.80)

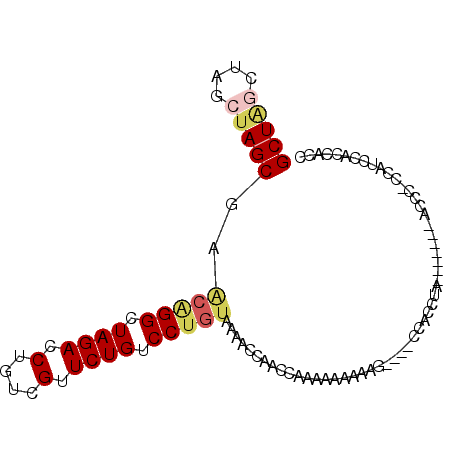
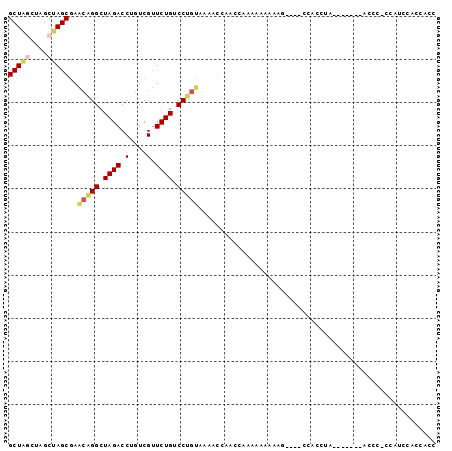
| Location | 15,117,959 – 15,118,077 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 84.02 |
| Mean single sequence MFE | -30.14 |
| Consensus MFE | -20.32 |
| Energy contribution | -21.60 |
| Covariance contribution | 1.28 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.67 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.513758 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15117959 118 + 22407834 GUGGGUAGGUGGUUUUUUUUUUUUUUGGUUGGUUUUACAGGACAGAACGACAGGUCUAGCCUGUUCGCUAGCUAGCUAGCUAGUUUGUGUGUUGCAGAAAUUUACUGUAAAAUACAAU .....(((.((((............((((((((...(((((..(((.(....).)))..)))))..)))))))))))).)))..((((((.((((((.......)))))).)))))). ( -32.20) >DroSec_CAF1 62748 104 + 1 ------AGGUUG----CUUUUUUUUUGGUUGGUUUUACAGGACAGAACGACAGGUCUAGCCUGCUCGCUAGC----UAGCUAGUUUGUGUGUUGCACAAAUUUACUGUAAAAUACAAU ------......----...........(((((((((((((.......(((((((.....)))).)))((((.----...))))((((((.....))))))....))))))))).)))) ( -27.00) >DroSim_CAF1 60435 104 + 1 ------AGGUUG----CUUUUUUUUUGGUUGGUUUUACAGGACAGAACGACAGGUCUAGCCUGUUCGCUAGC----UAGCUAGUUUGUGUGUUGCACAAAUUUACUGUAAAAUACAAU ------......----...........(((((((((((((........((((((.....)))))).(((((.----...)))))(((((.....))))).....))))))))).)))) ( -27.20) >DroEre_CAF1 63757 107 + 1 -----UGGGUG-----AUUUCCCUUUGGCUGG-UUUACAGGACAGAGCGACAGGUCUAGCCCGCUUGCUAGCUAGCCAGCUAGUUUGUGUGCUGCAGAAAUUUACUGUAAAAUACAAU -----.(((..-----....))).((((((((-((..(.((...(((((...((.....))))))).)).)..)))))))))).((((((..(((((.......)))))..)))))). ( -35.80) >DroYak_CAF1 77689 102 + 1 -----UAGGUG-----GUUU---UUUGGGUGGAUUUGCAGGACAGAACGACAGGUCUAGCC---UCGCUCGCUAGCUAGCUAGUUUGUGUGUUGCAGCAAUUUACUGUAAAAUACAAU -----(((.((-----((..---...((.((((((((..(.......)..)))))))).))---......)))).)))......((((((.((((((.......)))))).)))))). ( -28.50) >consensus _____UAGGUGG____CUUUUUUUUUGGUUGGUUUUACAGGACAGAACGACAGGUCUAGCCUGCUCGCUAGCUAGCUAGCUAGUUUGUGUGUUGCAGAAAUUUACUGUAAAAUACAAU ......................(((((.(((......)))..))))).((((((.....)))).))((((((......))))))((((((.((((((.......)))))).)))))). (-20.32 = -21.60 + 1.28)


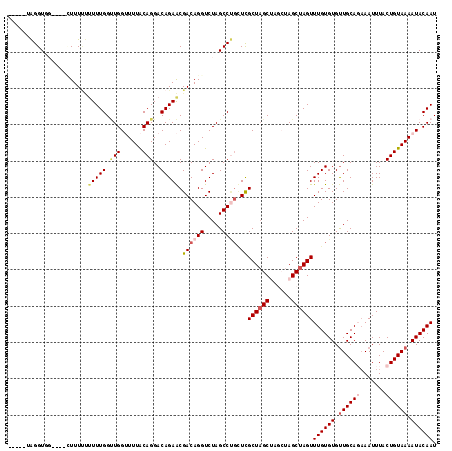
| Location | 15,117,959 – 15,118,077 |
|---|---|
| Length | 118 |
| Sequences | 5 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 84.02 |
| Mean single sequence MFE | -22.04 |
| Consensus MFE | -14.52 |
| Energy contribution | -15.04 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.79 |
| Structure conservation index | 0.66 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.578150 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 15117959 118 - 22407834 AUUGUAUUUUACAGUAAAUUUCUGCAACACACAAACUAGCUAGCUAGCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAAAAAACCACCUACCCAC ......((((((((.....................((((((....))))))(((.((((.....))))))).......))))))))................................ ( -20.80) >DroSec_CAF1 62748 104 - 1 AUUGUAUUUUACAGUAAAUUUGUGCAACACACAAACUAGCUA----GCUAGCGAGCAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG----CAACCU------ ......((((((((....((((((.....))))))((((((.----(((....))).))))))...............))))))))................----......------ ( -23.00) >DroSim_CAF1 60435 104 - 1 AUUGUAUUUUACAGUAAAUUUGUGCAACACACAAACUAGCUA----GCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG----CAACCU------ ......((((((((....((((((.....))))))((((...----.))))(((.((((.....))))))).......))))))))................----......------ ( -20.80) >DroEre_CAF1 63757 107 - 1 AUUGUAUUUUACAGUAAAUUUCUGCAGCACACAAACUAGCUGGCUAGCUAGCAAGCGGGCUAGACCUGUCGCUCUGUCCUGUAAA-CCAGCCAAAGGGAAAU-----CACCCA----- .......(((((((.........((((..((((..((((((.(((.(....).))).))))))...))).)..)))).)))))))-.........(((....-----..))).----- ( -26.20) >DroYak_CAF1 77689 102 - 1 AUUGUAUUUUACAGUAAAUUGCUGCAACACACAAACUAGCUAGCUAGCGAGCGA---GGCUAGACCUGUCGUUCUGUCCUGCAAAUCCACCCAAA---AAAC-----CACCUA----- ((((((...))))))...((((.(((..(((((..((((((.(((....)))..---))))))...))).))..)))...))))...........---....-----......----- ( -19.40) >consensus AUUGUAUUUUACAGUAAAUUUCUGCAACACACAAACUAGCUAGCUAGCUAGCGAACAGGCUAGACCUGUCGUUCUGUCCUGUAAAACCAACCAAAAAAAAAG____CCACCUA_____ ((((((...)))))).......((((..(((....(((((......)))))(((.((((.....)))))))...)))..))))................................... (-14.52 = -15.04 + 0.52)

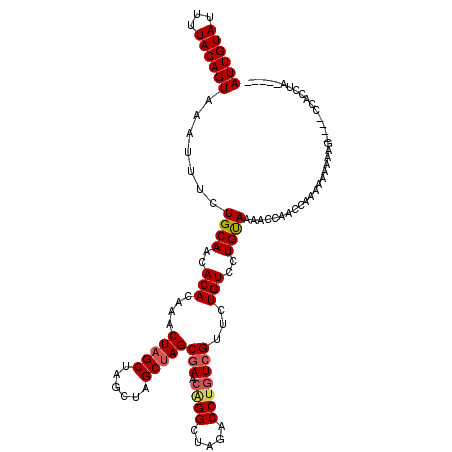

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:41:43 2006