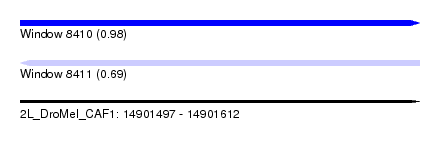

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,901,497 – 14,901,612 |
| Length | 115 |
| Max. P | 0.981560 |

| Location | 14,901,497 – 14,901,612 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.06 |
| Mean single sequence MFE | -37.12 |
| Consensus MFE | -32.28 |
| Energy contribution | -31.65 |
| Covariance contribution | -0.63 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.89 |
| SVM RNA-class probability | 0.981560 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
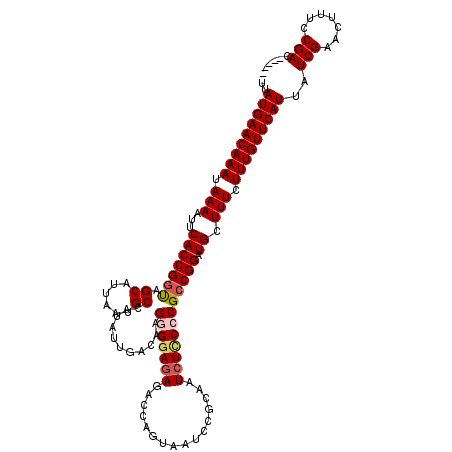
>2L_DroMel_CAF1 14901497 115 + 22407834 -----GUCGGAAAGUUCGAUAAUGAACAAAGAAAGCAUCCGGCGGGGAGAUUGCGGAUUACUGGUCUCUUCUCUUGUCAAUAUGGCAUUAAUGCUACCGGUGAAUUUUAUUUGUUCAUAA -----((((((...)))))).(((((((((((((....((((.((((((((..(((....)))))))))))...........(((((....)))))))))....)))).))))))))).. ( -33.30) >DroSec_CAF1 59075 115 + 1 -----CUCGGAAAGUUCGAUAAUGAACAAAGAAAGCAUCCGGCGGGGAGAUUGCGGAUUACUGGUCUCUCCCCUUGUCAAUAUGGCAUUAAUGCUAUCGGUGAAUUUUAUUUGUUCAUAA -----.(((((...)))))..(((((((((((((....(((..(((((((...(((....)))...)))))))........((((((....)))))))))....)))).))))))))).. ( -34.70) >DroSim_CAF1 61957 115 + 1 -----GUCGGAAAGUUCGAUAAUGAACAAAGAAAGCAUCCGGCGGGGAGAUUGCGGAUUACUGGUCUCUCCCCUUGUCAAUAUGGCAUUAAUGCUACCGGUGAAUUUUAUUUGUUCAUAA -----((((((...)))))).(((((((((((((....((((.(((((((...(((....)))...))))))).........(((((....)))))))))....)))).))))))))).. ( -38.30) >DroYak_CAF1 58769 120 + 1 GUAGCGUCGGAAAGUUCGAUAAUGAACAAAGAAAGCAUCCGGCGGGAAGAUUGGGGAUUACUGCUCUUUCCGCUCGUCAAUAUGGCCUUAAUGCUGCCGGUGAGCUUUAUUUGUUCAUAA .....((((((...)))))).(((((((((.(((((...(((((((((((..((......))..))))))))).))(((...((((.........)))).)))))))).))))))))).. ( -42.20) >consensus _____GUCGGAAAGUUCGAUAAUGAACAAAGAAAGCAUCCGGCGGGGAGAUUGCGGAUUACUGGUCUCUCCCCUUGUCAAUAUGGCAUUAAUGCUACCGGUGAAUUUUAUUUGUUCAUAA .....((((((...)))))).(((((((((.((((...((((.((((((((((.(.....)))))))))))...........((((......))))))))....)))).))))))))).. (-32.28 = -31.65 + -0.63)



| Location | 14,901,497 – 14,901,612 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.06 |
| Mean single sequence MFE | -27.96 |
| Consensus MFE | -22.90 |
| Energy contribution | -23.84 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.692266 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14901497 115 - 22407834 UUAUGAACAAAUAAAAUUCACCGGUAGCAUUAAUGCCAUAUUGACAAGAGAAGAGACCAGUAAUCCGCAAUCUCCCCGCCGGAUGCUUUCUUUGUUCAUUAUCGAACUUUCCGAC----- ..(((((((((.(((...(((((((.(((....)))................((((...((.....))..))))...))))).)).))).)))))))))..(((.......))).----- ( -21.00) >DroSec_CAF1 59075 115 - 1 UUAUGAACAAAUAAAAUUCACCGAUAGCAUUAAUGCCAUAUUGACAAGGGGAGAGACCAGUAAUCCGCAAUCUCCCCGCCGGAUGCUUUCUUUGUUCAUUAUCGAACUUUCCGAG----- ..(((((((((.(((((((..((((((((....)))..)))))....(((((((...(........)...)))))))...))))..))).)))))))))..(((.......))).----- ( -27.40) >DroSim_CAF1 61957 115 - 1 UUAUGAACAAAUAAAAUUCACCGGUAGCAUUAAUGCCAUAUUGACAAGGGGAGAGACCAGUAAUCCGCAAUCUCCCCGCCGGAUGCUUUCUUUGUUCAUUAUCGAACUUUCCGAC----- ..(((((((((.(((...(((((((.(((....)))...........(((((((...(........)...)))))))))))).)).))).)))))))))..(((.......))).----- ( -31.90) >DroYak_CAF1 58769 120 - 1 UUAUGAACAAAUAAAGCUCACCGGCAGCAUUAAGGCCAUAUUGACGAGCGGAAAGAGCAGUAAUCCCCAAUCUUCCCGCCGGAUGCUUUCUUUGUUCAUUAUCGAACUUUCCGACGCUAC ..(((((((((.(((((((...(((.........))).......((.((((.((((..............)))).)))))))).))))).)))))))))..(((.......)))...... ( -31.54) >consensus UUAUGAACAAAUAAAAUUCACCGGUAGCAUUAAUGCCAUAUUGACAAGGGGAGAGACCAGUAAUCCGCAAUCUCCCCGCCGGAUGCUUUCUUUGUUCAUUAUCGAACUUUCCGAC_____ ..(((((((((.(((...(((((((.((......))...........(((((((................)))))))))))).)).))).)))))))))..(((.......)))...... (-22.90 = -23.84 + 0.94)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:36 2006