
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,900,278 – 14,900,396 |
| Length | 118 |
| Max. P | 0.654774 |

| Location | 14,900,278 – 14,900,396 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 91.27 |
| Mean single sequence MFE | -30.55 |
| Consensus MFE | -23.88 |
| Energy contribution | -25.00 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.554474 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14900278 118 + 22407834 ACAUAUAUACGUUUAUUCGUUUAGGCCAAACACAAGCGCGCUUAGCCAUAAAUUGUUGCGUUGAGCGACGCACGGAAAAGCGGAGAACGGCAUAAAA-AAAAGUAGGAGCAGCUCCGGG .........(((((.(((((((.(......)....(((((((((((((........)).)))))))).)))......))))))))))))........-.......((((...))))... ( -31.10) >DroSec_CAF1 57884 119 + 1 AAAAAUAUACGUUUAUACUUUAAGGCAAAACACAAGCGCGCUUAGCCAUAAAUUGUUGCGUUGAGCGACGCACGGAAAAGCGAAGAACGGCAUAAAAAAAAAGUUGGAGCAGCUCCGGG .........(((((.....................(((((((((((((........)).)))))))).))).((......))..)))))..............((((((...)))))). ( -26.70) >DroSim_CAF1 60725 117 + 1 AUAAAUAUACGUUUAUACUUUUAGGCCAAACACAAGCGCGCUUAGCCAUAAAUUGUUGCGUUGAGCGACGCACGGAAAAGCGAAGAACGGCAUAAAA--AAAGUAGGAGCAGCUCCGGG ..........((((.(((((((..(((........(((((((((((((........)).)))))))).))).((......))......))).....)--)))))).))))......... ( -31.50) >DroYak_CAF1 57525 113 + 1 AAAUAUAUACGCUCAUACUUUUAGGCCAAACACAAGC--GCUUAGCCAUAAAUUGUUGCGUUGAGCGACGCACGGAAAAGCGGAGAGCGGCAUAAA---AAAGUAGGAGCCGCUC-GGA ..........((((.(((((((..(((.........(--(((((((((........)).)))))))).(((........)))......)))....)---)))))).)))).....-... ( -32.90) >consensus AAAAAUAUACGUUUAUACUUUUAGGCCAAACACAAGCGCGCUUAGCCAUAAAUUGUUGCGUUGAGCGACGCACGGAAAAGCGAAGAACGGCAUAAAA__AAAGUAGGAGCAGCUCCGGG ..........((((.((((((...(((........(((((((((((((........)).)))))))).))).((......))......)))........)))))).))))......... (-23.88 = -25.00 + 1.12)

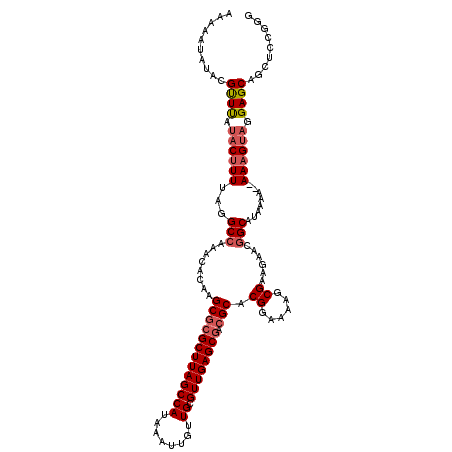

| Location | 14,900,278 – 14,900,396 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 91.27 |
| Mean single sequence MFE | -30.11 |
| Consensus MFE | -23.98 |
| Energy contribution | -25.35 |
| Covariance contribution | 1.37 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.654774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14900278 118 - 22407834 CCCGGAGCUGCUCCUACUUUU-UUUUAUGCCGUUCUCCGCUUUUCCGUGCGUCGCUCAACGCAACAAUUUAUGGCUAAGCGCGCUUGUGUUUGGCCUAAACGAAUAAACGUAUAUAUGU ...((((...)))).......-...(((((.((((............(((((......)))))....((((.((((((((((....)))))))))))))).))))....)))))..... ( -27.70) >DroSec_CAF1 57884 119 - 1 CCCGGAGCUGCUCCAACUUUUUUUUUAUGCCGUUCUUCGCUUUUCCGUGCGUCGCUCAACGCAACAAUUUAUGGCUAAGCGCGCUUGUGUUUUGCCUUAAAGUAUAAACGUAUAUUUUU ...((((...)))).........(((((((.................(((((......)))))....((((.(((.((((((....)))))).))).)))))))))))........... ( -25.40) >DroSim_CAF1 60725 117 - 1 CCCGGAGCUGCUCCUACUUU--UUUUAUGCCGUUCUUCGCUUUUCCGUGCGUCGCUCAACGCAACAAUUUAUGGCUAAGCGCGCUUGUGUUUGGCCUAAAAGUAUAAACGUAUAUUUAU ...((((...))))......--...(((((.(((..(.((((((...(((((......))))).........((((((((((....)))))))))).)))))))..))))))))..... ( -29.80) >DroYak_CAF1 57525 113 - 1 UCC-GAGCGGCUCCUACUUU---UUUAUGCCGCUCUCCGCUUUUCCGUGCGUCGCUCAACGCAACAAUUUAUGGCUAAGC--GCUUGUGUUUGGCCUAAAAGUAUGAGCGUAUAUAUUU ...-(((((((.........---.....)))))))..(((((...(.(((((......)))))....((((.((((((((--(....))))))))))))).)...)))))......... ( -37.54) >consensus CCCGGAGCUGCUCCUACUUU__UUUUAUGCCGUUCUCCGCUUUUCCGUGCGUCGCUCAACGCAACAAUUUAUGGCUAAGCGCGCUUGUGUUUGGCCUAAAAGUAUAAACGUAUAUAUUU ...((((...))))...........(((((.(((....((((((...(((((......))))).........((((((((((....)))))))))).))))))...))))))))..... (-23.98 = -25.35 + 1.37)


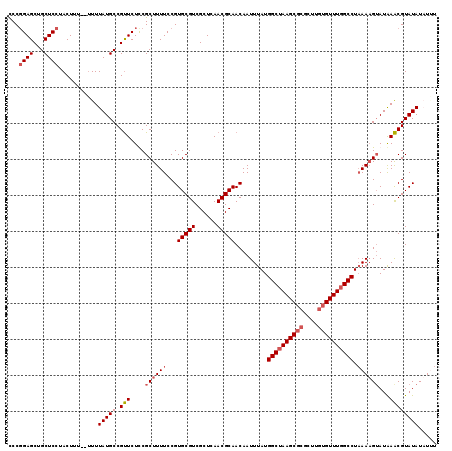
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:33 2006