
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,895,046 – 14,895,141 |
| Length | 95 |
| Max. P | 0.828334 |

| Location | 14,895,046 – 14,895,141 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 80.02 |
| Mean single sequence MFE | -33.60 |
| Consensus MFE | -20.04 |
| Energy contribution | -19.41 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.18 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.71 |
| SVM RNA-class probability | 0.828334 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
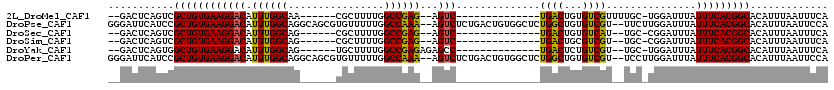
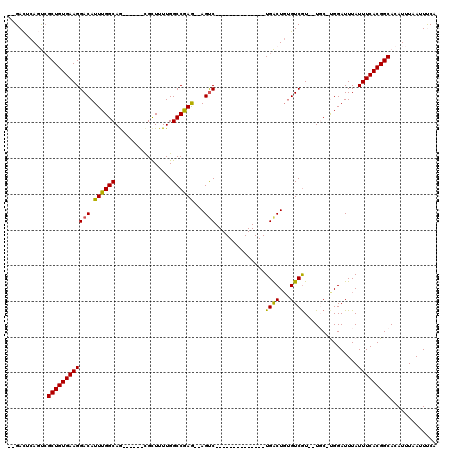
>2L_DroMel_CAF1 14895046 95 - 22407834 --GACUCAGUCGCUGUGAAGGACAUUUGGCAA------CGCUUUUGGCCGAG--AGUC--------------UGACUGUGUCGUUUUGC-UGGAUUUAUUUCACGGCACAUUUAAUUUCA --(((...)))(((((((((...(((..((((------(((..(..(.(...--.).)--------------..)..)))).....)))-..)))...)))))))))............. ( -28.70) >DroPse_CAF1 111064 116 - 1 GGGAUUCAUCCGCUGUGAAGGACAUUUGGCAGGCAGCGUGUUUUUGGCCAAA--AGUCUCUGACUGUGGCUCUGGCUGUGUCGU--UUCUUGGAUUUAUUUCACGGCACAUUUAAUUCCA ((((((.....((((((((((((.((((((.(((.....)))....))))))--.))))..(((.(..((....))..))))..--.............))))))))......)))))). ( -35.40) >DroSec_CAF1 52739 93 - 1 --GACUCAGUCGCUGUGAAGGACAUUUGGCAG------CGCUUUUGGCCGAG--AGUC--------------UGACUGUGUCAU--UGC-CGGAUUUAUUUCACGGCACAUUUAAUUUCA --(((.((((((((((.(((....))).))))------)((((((....)))--))).--------------.))))).)))..--(((-((((......)).)))))............ ( -34.10) >DroSim_CAF1 52586 93 - 1 --GACUCAGUCGCUGUGAAGGACAUUUGGCAG------CGCUUUUGGCCGAG--AGUC--------------UGACUGCGUCGU--UGC-CGGAUUUAUUUCACGGCACAUUUAAUUUCA --(((.((((((((((.(((....))).))))------)((((((....)))--))).--------------.))))).)))((--.((-((((......)).))))))........... ( -34.00) >DroYak_CAF1 52226 95 - 1 --GACUCAGUGGCUGUGAAGGACAUUUGGCAG------UGCUUUUGGCCGAGAGAGCC--------------UGACUCUGUCGU--UGC-UGGAUUUAUUUCACGGCACAUUUAAUUUCA --.....(((((((((((((...(((..((((------.((.....))(((.((((..--------------...)))).))))--)))-..)))...))))))))).))))........ ( -32.60) >DroPer_CAF1 93695 116 - 1 GGGAUUCAUCCGCUGUGAAGGACAUUUGGCAGGCAGCGUGUUUUUGGCCAAA--AGUCUCUGACUGUGGCUCUGGCUGUGUCGU--UCCUUGGAUUUAUUUCACGGCACAUUUAAUUCCA ((((((.....((((((((((((((..(.(((((.(((.(((...(((....--.)))...)))))).)).))).).)))))..--((....))....)))))))))......)))))). ( -36.80) >consensus __GACUCAGUCGCUGUGAAGGACAUUUGGCAG______CGCUUUUGGCCGAG__AGUC______________UGACUGUGUCGU__UGC_UGGAUUUAUUUCACGGCACAUUUAAUUUCA ...........((((((((((((.((((((................))))))...)))..............((((...))))...............)))))))))............. (-20.04 = -19.41 + -0.64)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:40:26 2006