
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,786,639 – 14,786,774 |
| Length | 135 |
| Max. P | 0.999774 |

| Location | 14,786,639 – 14,786,737 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 97.55 |
| Mean single sequence MFE | -30.72 |
| Consensus MFE | -28.90 |
| Energy contribution | -29.30 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.71 |
| Structure conservation index | 0.94 |
| SVM decision value | 4.05 |
| SVM RNA-class probability | 0.999774 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14786639 98 + 22407834 CAGUCCAGCGCACAACUUCACAGGUGCCAAAUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCG ...................(((((.((.(((.(((((((.....(((((((...((....))...)))))))...))))))).)))..)).))))).. ( -30.10) >DroSec_CAF1 33145 98 + 1 CAGUCCAGCGCACAACUUUACAGGUGCCAAAUGACAACAACAAAUUGCCCGAUUGCCUUCGCUACUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCG ...................(((((.((.(((.(((((((.....(((((((...((....))...)))))))...))))))).)))..)).))))).. ( -30.00) >DroSim_CAF1 31217 98 + 1 CAGUCCAGCGCACAACUUCACAGGUGCCAAAUGACAACAACAAAUUGCCCGAUUGCCUUCGCUACUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCG ...................(((((.((.(((.(((((((.....(((((((...((....))...)))))))...))))))).)))..)).))))).. ( -30.10) >DroEre_CAF1 25950 98 + 1 CAGUGCAGCGCACAACUUCACAGGUGCCAACUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCG ..((((...))))......(((((.((.....(((((((.....(((((((...((....))...)))))))...)))))))......)).))))).. ( -33.50) >DroYak_CAF1 30304 98 + 1 CAGUGCAGCGCACAACUUCACAGGUGCCAAAUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUCGUCCUUUCCGCUCCUGUCG ..((((...))))......(((((.((.(((.(((.(((.....(((((((...((....))...)))))))...))).))).)))..)).))))).. ( -29.90) >consensus CAGUCCAGCGCACAACUUCACAGGUGCCAAAUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCG ...................(((((.((.(((.(((((((.....(((((((...((....))...)))))))...))))))).)))..)).))))).. (-28.90 = -29.30 + 0.40)


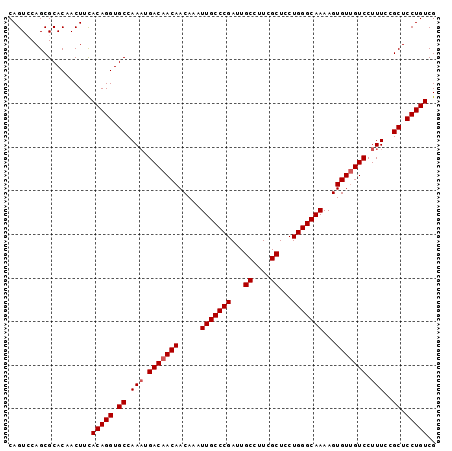
| Location | 14,786,639 – 14,786,737 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | reverse |
| Mean pairwise identity | 97.55 |
| Mean single sequence MFE | -38.08 |
| Consensus MFE | -35.32 |
| Energy contribution | -35.16 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.76 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.91 |
| SVM RNA-class probability | 0.999697 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14786639 98 - 22407834 CGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAUUUGGCACCUGUGAAGUUGUGCGCUGGACUG ..(((((.((.(((..((((((((..((((((.(..((....))..).))))))...).))))))).))).)).)))))..((((........)))). ( -36.60) >DroSec_CAF1 33145 98 - 1 CGACAGGAGCGGAAAGGACAACACUUUUGCCCAGUAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAUUUGGCACCUGUAAAGUUGUGCGCUGGACUG ..(((((.((.(((..((((((((..((((((.((.((....)).)).))))))...).))))))).))).)).)))))..((((........)))). ( -37.70) >DroSim_CAF1 31217 98 - 1 CGACAGGAGCGGAAAGGACAACACUUUUGCCCAGUAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAUUUGGCACCUGUGAAGUUGUGCGCUGGACUG ..(((((.((.(((..((((((((..((((((.((.((....)).)).))))))...).))))))).))).)).)))))..((((........)))). ( -37.70) >DroEre_CAF1 25950 98 - 1 CGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAGUUGGCACCUGUGAAGUUGUGCGCUGCACUG ..(((((.((.((...((((((((..((((((.(..((....))..).))))))...).)))))))..)).)).)))))......((((...)))).. ( -39.70) >DroYak_CAF1 30304 98 - 1 CGACAGGAGCGGAAAGGACGACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAUUUGGCACCUGUGAAGUUGUGCGCUGCACUG ..(((((.((.(((..((((((((..((((((.(..((....))..).))))))...).))))))).))).)).)))))......((((...)))).. ( -38.70) >consensus CGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAUUUGGCACCUGUGAAGUUGUGCGCUGGACUG ..(((((.((.((...((((((((..((((((....((....))....))))))...).)))))))..)).)).)))))................... (-35.32 = -35.16 + -0.16)
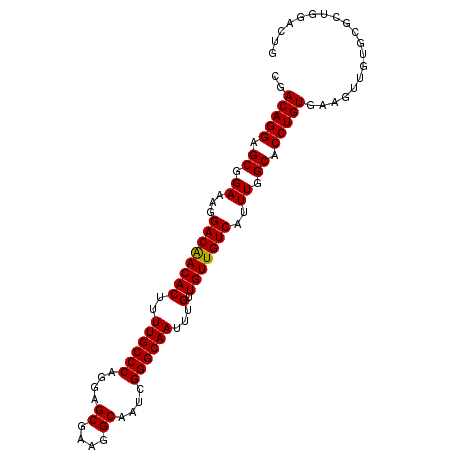


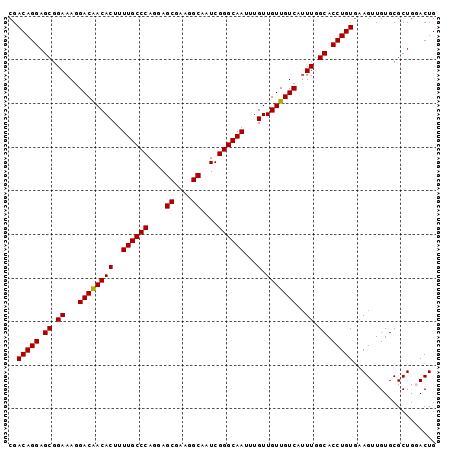
| Location | 14,786,669 – 14,786,774 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 96.00 |
| Mean single sequence MFE | -33.57 |
| Consensus MFE | -31.49 |
| Energy contribution | -32.18 |
| Covariance contribution | 0.70 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.94 |
| SVM decision value | 2.72 |
| SVM RNA-class probability | 0.996604 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14786669 105 + 22407834 AUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..(((((((.....(((((((...((....))...)))))))...))))))).((((((....(((((((......)))))))..))))))(((........))) ( -34.80) >DroSec_CAF1 33175 105 + 1 AUGACAACAACAAAUUGCCCGAUUGCCUUCGCUACUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..(((((((.....(((((((...((....))...)))))))...))))))).((((((....(((((((......)))))))..))))))(((........))) ( -34.80) >DroSim_CAF1 31247 105 + 1 AUGACAACAACAAAUUGCCCGAUUGCCUUCGCUACUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..(((((((.....(((((((...((....))...)))))))...))))))).((((((....(((((((......)))))))..))))))(((........))) ( -34.80) >DroEre_CAF1 25980 105 + 1 CUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..(((((((.....(((((((...((....))...)))))))...))))))).((((((....(((((((......)))))))..))))))(((........))) ( -34.80) >DroYak_CAF1 30334 105 + 1 AUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUCGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..((((........(((((((...((....))...)))))))...))))((((..((.((((((((((((......)))))))...)))...)).)).))))... ( -31.60) >DroAna_CAF1 22829 105 + 1 AUGACAACAACAAAUUGCCCGAUUGCCUCCGCUCCAGGGCAAGAGGGUUGUCCUUGCCGAUGCUGUCGAUGACGAUGUCGACAUAUGGAAAUACUGAUGGAUGCG ..((((((......((((((....((....))....))))))....))))))....((.(((.(((((((......))))))))))))................. ( -30.60) >consensus AUGACAACAACAAAUUGCCCGAUUGCCUUCGCUCCUGGGCAAAAGUGUUGUCCUUUCCGCUCCUGUCGAUGACGAUGUCGACAUAUGGAAAUGCUGAUGGAUGCA ..(((((((.....(((((((...((....))...)))))))...))))))).((((((....(((((((......)))))))..))))))(((........))) (-31.49 = -32.18 + 0.70)


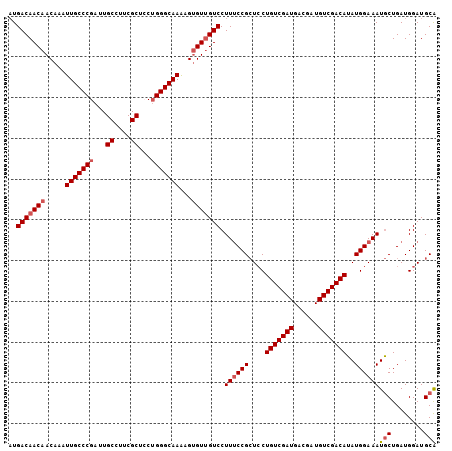
| Location | 14,786,669 – 14,786,774 |
|---|---|
| Length | 105 |
| Sequences | 6 |
| Columns | 105 |
| Reading direction | reverse |
| Mean pairwise identity | 96.00 |
| Mean single sequence MFE | -35.52 |
| Consensus MFE | -33.03 |
| Energy contribution | -33.25 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.93 |
| SVM decision value | 3.61 |
| SVM RNA-class probability | 0.999449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14786669 105 - 22407834 UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAU (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.(..((....))..).))))))...).))))))).. ( -36.10) >DroSec_CAF1 33175 105 - 1 UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACAACACUUUUGCCCAGUAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAU (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.((.((....)).)).))))))...).))))))).. ( -37.20) >DroSim_CAF1 31247 105 - 1 UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACAACACUUUUGCCCAGUAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAU (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.((.((....)).)).))))))...).))))))).. ( -37.20) >DroEre_CAF1 25980 105 - 1 UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAG (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.(..((....))..).))))))...).))))))).. ( -36.10) >DroYak_CAF1 30334 105 - 1 UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACGACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAU (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.(..((....))..).))))))...).))))))).. ( -35.80) >DroAna_CAF1 22829 105 - 1 CGCAUCCAUCAGUAUUUCCAUAUGUCGACAUCGUCAUCGACAGCAUCGGCAAGGACAACCCUCUUGCCCUGGAGCGGAGGCAAUCGGGCAAUUUGUUGUUGUCAU .((........((((....))))(((((........))))).))...(((((.((((((((..((((((((...))).)))))..)))....))))).))))).. ( -30.70) >consensus UGCAUCCAUCAGCAUUUCCAUAUGUCGACAUCGUCAUCGACAGGAGCGGAAAGGACAACACUUUUGCCCAGGAGCGAAGGCAAUCGGGCAAUUUGUUGUUGUCAU (((........)))(((((...((((((........)))))).....))))).((((((((..((((((.(..((....))..).))))))...).))))))).. (-33.03 = -33.25 + 0.22)


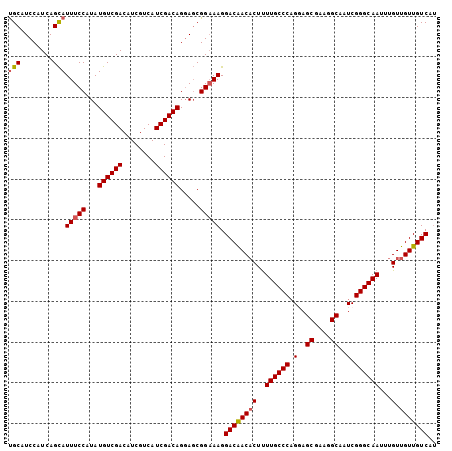
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:39:27 2006