
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,702,736 – 14,702,828 |
| Length | 92 |
| Max. P | 0.997121 |

| Location | 14,702,736 – 14,702,828 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 71.19 |
| Mean single sequence MFE | -24.82 |
| Consensus MFE | -3.98 |
| Energy contribution | -6.78 |
| Covariance contribution | 2.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.16 |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.600636 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
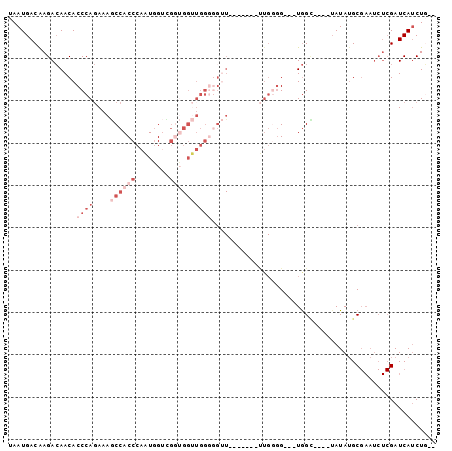
>2L_DroMel_CAF1 14702736 92 + 22407834 UAAUGACAAGACAACACCCAGAAAGCCACCCAAUGGUUGGUGGUUGGGGGUUUGGGGUUUUGGGG---UGGU----UAUAUGCGAAUCUCGAUCAUCUU-- ...........(((.((((...(((((.((((((.(....).))))))))))).)))).)))(((---((((----(.............)))))))).-- ( -28.52) >DroSec_CAF1 58896 84 + 1 UAAUGACAAGACAACACCCAGAAAGCCACCCAAUGGUCGGUGGUUGGGGGUU-------UUG-GG---UGGC----UAUAUGCGAAUCUCGAUCAUCUG-- ..(((((.(((...((((((((..(((.((((((.(....).))))))))))-------)))-))---))((----.....))...))).).))))...-- ( -28.90) >DroSim_CAF1 59173 78 + 1 UAAUGACAAGACAACACCCAGAAAGCCACCCAAUGGUCGGUGGUUGGGGGUU-------UUGGGG---UGGC----UAUAAGC-------GAUCAUCUG-- ..(((((.......(((((..((((((.((((((.(....).))))))))))-------)).)))---))((----.....))-------).))))...-- ( -27.50) >DroEre_CAF1 58473 76 + 1 UAAUGACAAGACAACACCCAGAAAACCACCCAAUGGAUA-------CCAA-------------UG---UGGUAUGGGAUAUGCGAAUCUCGAUCAACUG-- ...(((..........((((....(((((....(((...-------))).-------------.)---)))).)))).....((.....)).)))....-- ( -13.50) >DroAna_CAF1 55941 87 + 1 UAAUGACGAGAGAACACCCA-AAAACCACCCAGUUGGGGCGGGCUGG--GCU-------CUGGGGAUAUAGA----UAUAUGCGAAUCUCGAUCAUCUCGG ......(((((.....((((-.(..((((((.((....))))).)))--..)-------.))))(((..(((----(........))))..))).))))). ( -25.70) >consensus UAAUGACAAGACAACACCCAGAAAGCCACCCAAUGGUCGGUGGUUGGGGGUU_______UUGGGG___UGGC____UAUAUGCGAAUCUCGAUCAUCUG__ ................((((....((((((........))))))))))..................................................... ( -3.98 = -6.78 + 2.80)

| Location | 14,702,736 – 14,702,828 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 71.19 |
| Mean single sequence MFE | -24.79 |
| Consensus MFE | -9.93 |
| Energy contribution | -12.05 |
| Covariance contribution | 2.12 |
| Combinations/Pair | 1.17 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.40 |
| SVM decision value | 2.80 |
| SVM RNA-class probability | 0.997121 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
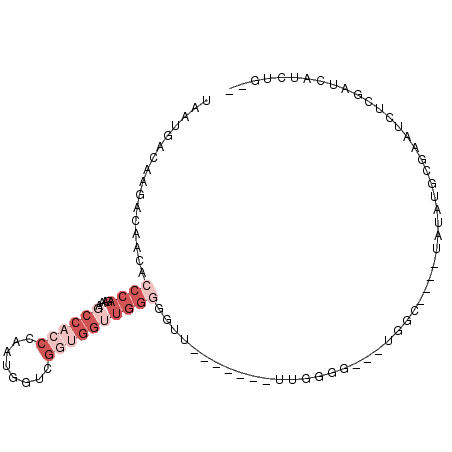
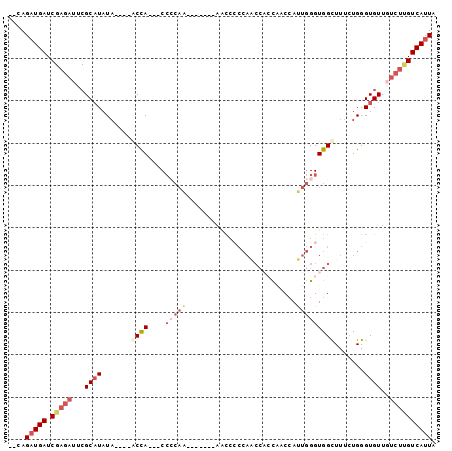
>2L_DroMel_CAF1 14702736 92 - 22407834 --AAGAUGAUCGAGAUUCGCAUAUA----ACCA---CCCCAAAACCCCAAACCCCCAACCACCAACCAUUGGGUGGCUUUCUGGGUGUUGUCUUGUCAUUA --..(((((.((((((..(((....----....---.((((.((..(((....(((((..........))))))))..)).))))))).))))))))))). ( -21.30) >DroSec_CAF1 58896 84 - 1 --CAGAUGAUCGAGAUUCGCAUAUA----GCCA---CC-CAA-------AACCCCCAACCACCGACCAUUGGGUGGCUUUCUGGGUGUUGUCUUGUCAUUA --..(((((.((((((.(((....(----((((---((-(((-------...................))))))))))......)))..))))))))))). ( -29.71) >DroSim_CAF1 59173 78 - 1 --CAGAUGAUC-------GCUUAUA----GCCA---CCCCAA-------AACCCCCAACCACCGACCAUUGGGUGGCUUUCUGGGUGUUGUCUUGUCAUUA --.(((..(.(-------(((((.(----((((---(((...-------.....................))))))))...)))))))..)))........ ( -22.46) >DroEre_CAF1 58473 76 - 1 --CAGUUGAUCGAGAUUCGCAUAUCCCAUACCA---CA-------------UUGG-------UAUCCAUUGGGUGGUUUUCUGGGUGUUGUCUUGUCAUUA --....(((.((((((..(((...((((.((((---(.-------------.(((-------...)))....)))))....))))))).)))))))))... ( -21.80) >DroAna_CAF1 55941 87 - 1 CCGAGAUGAUCGAGAUUCGCAUAUA----UCUAUAUCCCCAG-------AGC--CCAGCCCGCCCCAACUGGGUGGUUUU-UGGGUGUUCUCUCGUCAUUA ....(((((.(((((...((.....----(((........))-------)((--((((.((((((.....))))))...)-)))))))..)))))))))). ( -28.70) >consensus __CAGAUGAUCGAGAUUCGCAUAUA____ACCA___CCCCAA_______AACCCCCAACCACCAACCAUUGGGUGGCUUUCUGGGUGUUGUCUUGUCAUUA ....(((((.(((((...((((.......((((....(((((..........................))))))))).......))))..)))))))))). ( -9.93 = -12.05 + 2.12)
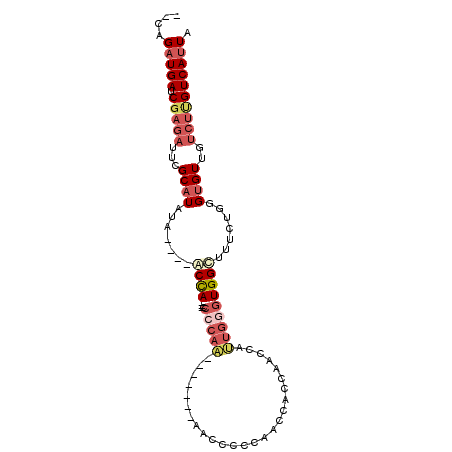


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:38:19 2006