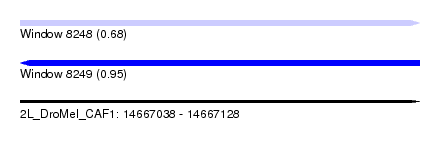

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,667,038 – 14,667,128 |
| Length | 90 |
| Max. P | 0.952493 |

| Location | 14,667,038 – 14,667,128 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 85.20 |
| Mean single sequence MFE | -15.56 |
| Consensus MFE | -11.12 |
| Energy contribution | -11.72 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.681726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
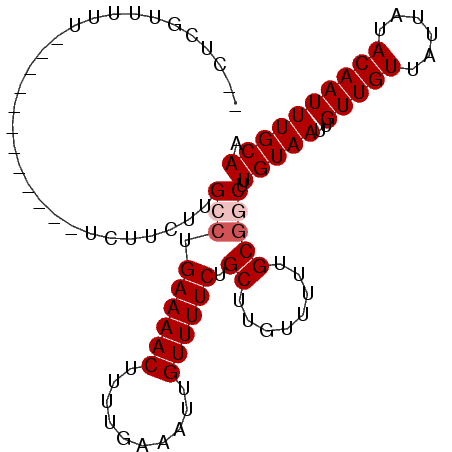
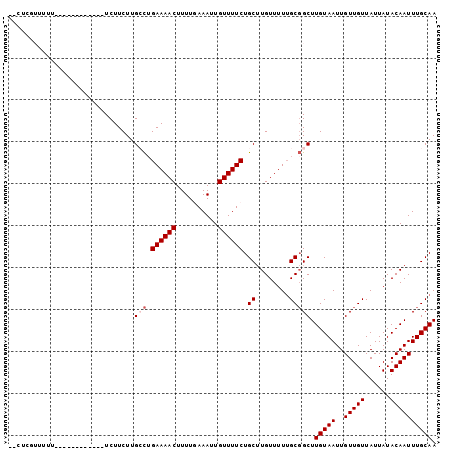
>2L_DroMel_CAF1 14667038 90 + 22407834 AACUCGUUUUU------------UCUUCUUGGCUGAAAACUUUUGAAAUUGUUUUCUGCUUGUUUUUGCGGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA ...((((....------------.(.....(((.((((((..........)))))).))).).....))))..(((((..(((((......)))))))))). ( -14.50) >DroSec_CAF1 21602 81 + 1 ---------UU------------UCUUCUUGCCUGAAAACUUUUGAAAUUGUUUUCUGCUUGUUUUUGCGGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA ---------..------------.......((((((((((....((((....)))).....))))))).))).(((((..(((((......)))))))))). ( -15.50) >DroSim_CAF1 23276 81 + 1 ---------UU------------UCUUCUUGCCUGAAAACUUUUGAAAUUGUUUUCUGCUUGUUUUUGCGGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA ---------..------------.......((((((((((....((((....)))).....))))))).))).(((((..(((((......)))))))))). ( -15.50) >DroEre_CAF1 21667 102 + 1 UGCUCGUUUUUGUCUCUUGUUUGUCUUCUUGGCAGAAAACUUUUAAAAUUGUUUUCUGCAUGUUUUUGCGGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA .((.(((....(.(........).)......(((((((((..........)))))))))........))))).(((((..(((((......)))))))))). ( -19.80) >DroYak_CAF1 22026 90 + 1 UUCUCGUUUUU------------UCUUCUUGCUUGAAAACUUUUAAAAUUGUUUUCUGCUUGUUUUGGCUGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA ...........------------.......((..((((((..........)))))).(((......))).)).(((((..(((((......)))))))))). ( -12.50) >consensus __CUCGUUUUU____________UCUUCUUGCCUGAAAACUUUUGAAAUUGUUUUCUGCUUGUUUUUGCGGCUUGUAAUUGUUGUUAUUAUACAAUUUGCAA ..............................(((.((((((..........)))))).((........))))).(((((..(((((......)))))))))). (-11.12 = -11.72 + 0.60)

| Location | 14,667,038 – 14,667,128 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 85.20 |
| Mean single sequence MFE | -12.97 |
| Consensus MFE | -9.22 |
| Energy contribution | -9.42 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.62 |
| Structure conservation index | 0.71 |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.952493 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
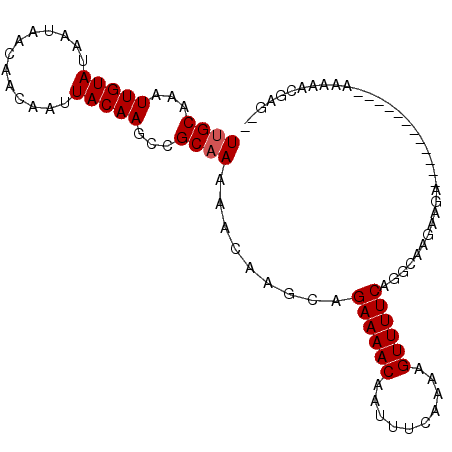
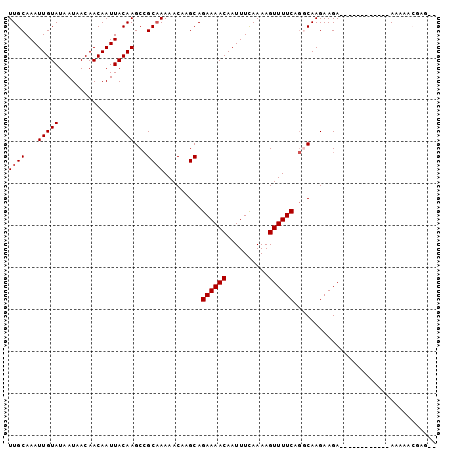
>2L_DroMel_CAF1 14667038 90 - 22407834 UUGCAAAUUGUAUAAUAACAACAAUUACAAGCCGCAAAAACAAGCAGAAAACAAUUUCAAAAGUUUUCAGCCAAGAAGA------------AAAAACGAGUU ((((...(((((.............)))))...))))......((.((((((..........)))))).))........------------........... ( -11.52) >DroSec_CAF1 21602 81 - 1 UUGCAAAUUGUAUAAUAACAACAAUUACAAGCCGCAAAAACAAGCAGAAAACAAUUUCAAAAGUUUUCAGGCAAGAAGA------------AA--------- .....((((((.........))))))....(((((........)).((((((..........)))))).))).......------------..--------- ( -13.20) >DroSim_CAF1 23276 81 - 1 UUGCAAAUUGUAUAAUAACAACAAUUACAAGCCGCAAAAACAAGCAGAAAACAAUUUCAAAAGUUUUCAGGCAAGAAGA------------AA--------- .....((((((.........))))))....(((((........)).((((((..........)))))).))).......------------..--------- ( -13.20) >DroEre_CAF1 21667 102 - 1 UUGCAAAUUGUAUAAUAACAACAAUUACAAGCCGCAAAAACAUGCAGAAAACAAUUUUAAAAGUUUUCUGCCAAGAAGACAAACAAGAGACAAAAACGAGCA ((((...(((((.............)))))...))))......(((((((((..........)))))))))............................... ( -16.62) >DroYak_CAF1 22026 90 - 1 UUGCAAAUUGUAUAAUAACAACAAUUACAAGCAGCCAAAACAAGCAGAAAACAAUUUUAAAAGUUUUCAAGCAAGAAGA------------AAAAACGAGAA ((((.((((((.........)))))).......((........)).((((((..........))))))..)))).....------------........... ( -10.30) >consensus UUGCAAAUUGUAUAAUAACAACAAUUACAAGCCGCAAAAACAAGCAGAAAACAAUUUCAAAAGUUUUCAGGCAAGAAGA____________AAAAACGAG__ ((((...(((((.............)))))...)))).........((((((..........)))))).................................. ( -9.22 = -9.42 + 0.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:38:01 2006