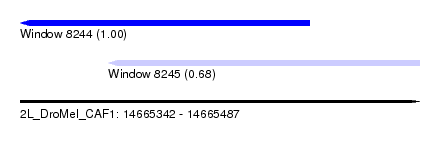

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,665,342 – 14,665,487 |
| Length | 145 |
| Max. P | 0.996804 |

| Location | 14,665,342 – 14,665,447 |
|---|---|
| Length | 105 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 82.64 |
| Mean single sequence MFE | -26.45 |
| Consensus MFE | -18.40 |
| Energy contribution | -19.90 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.07 |
| Mean z-score | -3.34 |
| Structure conservation index | 0.70 |
| SVM decision value | 2.75 |
| SVM RNA-class probability | 0.996804 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
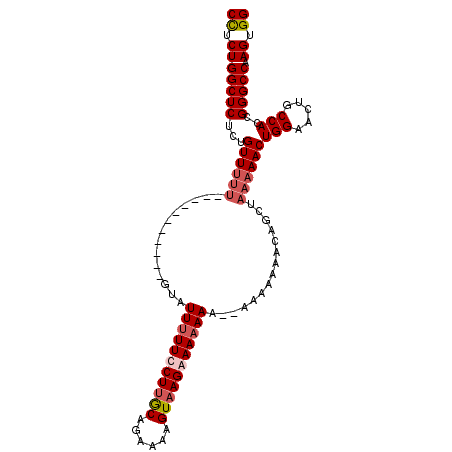
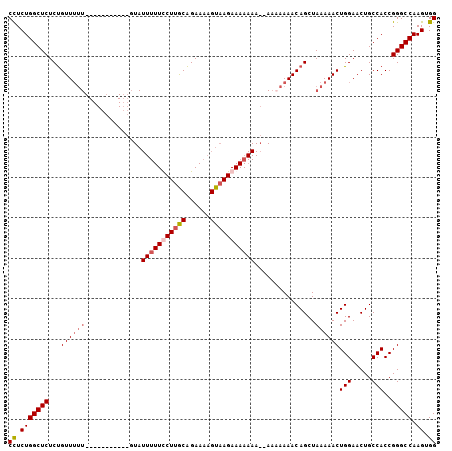
>2L_DroMel_CAF1 14665342 105 - 22407834 CCUCUGGCUCUCUGUUUUUUUUUUUCUGUUUUAUUUUUCCUUACAGAAAAGUAAGAAAAAAAA-AACAAAACAGCUAAAAACUGGAACUGCCACCGGGCCAAGUGG ((.(((((((.(((((((.((((((...(((((((((((......)))))))))))...))))-)).)))))))........(((.....)))..))))).)).)) ( -29.30) >DroSec_CAF1 19950 93 - 1 CCUCUGGCUCUCUGUUUUU-----------GUAUUUUUUCUUGCAGAAAAGUAAGAAAAAAA--AAAAAAACAGCUAAAAACUGGAACUGCCACCGGGCCAAGUGG ((.(((((((.((((((((-----------...(((((((((((......))))))))))).--..))))))))........(((.....)))..))))).)).)) ( -30.10) >DroSim_CAF1 21620 95 - 1 CCUCUGGCUCUCUGUUUUU-----------GUAUUUUUUCUUGCAGAAAAGUAAGAAAAAAAAAAAAAAAACAGCUAAAAACUGGAACUGCCACCGGGCCAAGUGG ((.(((((((.((((((((-----------.(.(((((((((((......))))))))))).)...))))))))........(((.....)))..))))).)).)) ( -28.80) >DroEre_CAF1 20007 80 - 1 CUUCUGGCUCUCCGUU--------------------UUCCUGGCGGAAAAGUAAGAAAA------ACAAAACAGCUAAAAACUGGAACUGCCACCGGGCCAAGUGG ....((((((((((((--------------------.....))))))............------......(((.......)))...........))))))..... ( -17.60) >consensus CCUCUGGCUCUCUGUUUUU___________GUAUUUUUCCUUGCAGAAAAGUAAGAAAAAAA__AAAAAAACAGCUAAAAACUGGAACUGCCACCGGGCCAAGUGG ((.(((((((...((((((..............(((((((((((......)))))))))))...............))))))(((.....)))..))))).)).)) (-18.40 = -19.90 + 1.50)

| Location | 14,665,374 – 14,665,487 |
|---|---|
| Length | 113 |
| Sequences | 5 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 82.32 |
| Mean single sequence MFE | -25.28 |
| Consensus MFE | -17.56 |
| Energy contribution | -17.88 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.677765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
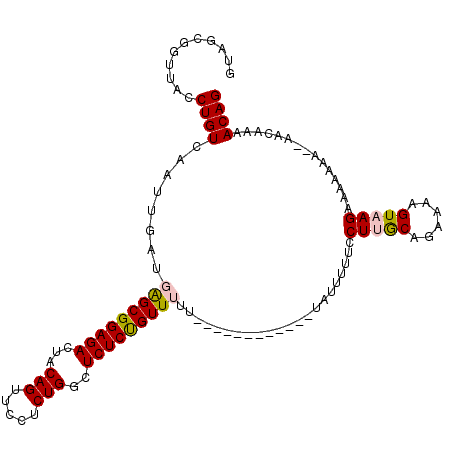
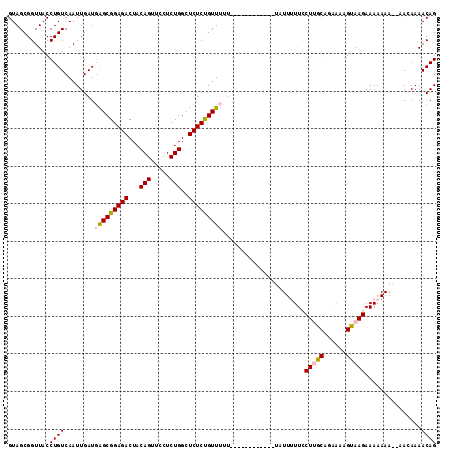
>2L_DroMel_CAF1 14665374 113 - 22407834 GUAGCGGUUACCUGUCAAUUGAUGAGCGGAGACUACAGUUCCUCUGGCUCUCUGUUUUUUUUUUUCUGUUUUAUUUUUCCUUACAGAAAAGUAAGAAAAAAAA-AACAAAACAG ...((((....))))........(((((....)..(((.....))))))).(((((((.((((((...(((((((((((......)))))))))))...))))-)).))))))) ( -27.20) >DroSec_CAF1 19982 101 - 1 GUAGCGGUUACCUGUCAAUUGAUGAGCGGAGACUACAGUUCCUCUGGCUCUCUGUUUUU-----------GUAUUUUUUCUUGCAGAAAAGUAAGAAAAAAA--AAAAAAACAG ...((((....))))........(((((....)..(((.....))))))).((((((((-----------...(((((((((((......))))))))))).--..)))))))) ( -28.00) >DroSim_CAF1 21652 103 - 1 GUAGCGGUUACCUGUCAAUUGAUGAGCGGAGACUACAGUUCCUCUGGCUCUCUGUUUUU-----------GUAUUUUUUCUUGCAGAAAAGUAAGAAAAAAAAAAAAAAAACAG ...((((....))))........(((((....)..(((.....))))))).((((((((-----------.(.(((((((((((......))))))))))).)...)))))))) ( -26.70) >DroEre_CAF1 20039 88 - 1 GUAGCGGUUACCUGUCAAUUGAUCGGCGGAGACUACAGUUCUUCUGGCUCUCCGUU--------------------UUCCUGGCGGAAAAGUAAGAAAA------ACAAAACAG (((((..(..((.(((....))).))..).).)))).(((.((((.(((.((((((--------------------.....))))))..))).)))).)------))....... ( -22.90) >DroYak_CAF1 20451 88 - 1 GUAGCGGUUACCUGUCAAUUGAUGAGCGGAGACUACAGUUCUUCUGGCUCUCUGUU--------------------UUCCUGGCAGUGAAGUAAGAAAA------ACAAAACAG ...((..((((.(((((...((.(((((((((...(((.....)))..))))))))--------------------))).))))))))).)).......------......... ( -21.60) >consensus GUAGCGGUUACCUGUCAAUUGAUGAGCGGAGACUACAGUUCCUCUGGCUCUCUGUUUUU____________UAUUUUUCCUUGCAGAAAAGUAAGAAAAAAA__AACAAAACAG ...........((((........(((((((((...(((.....)))..)))))))))......................(((((......)))))...............)))) (-17.56 = -17.88 + 0.32)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:37:58 2006