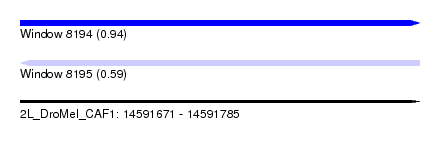

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,591,671 – 14,591,785 |
| Length | 114 |
| Max. P | 0.938415 |

| Location | 14,591,671 – 14,591,785 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.37 |
| Mean single sequence MFE | -34.20 |
| Consensus MFE | -29.47 |
| Energy contribution | -30.03 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.96 |
| Structure conservation index | 0.86 |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.938415 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
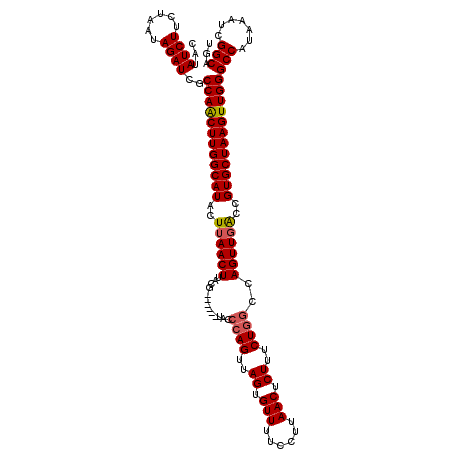
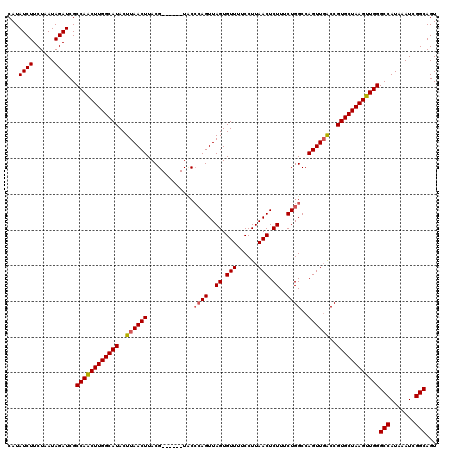
>2L_DroMel_CAF1 14591671 114 + 22407834 CAUAUCUUCUAAUAGAUCGCCAACUUGGCAUACUUAACUUACG------UUCGCAGUUAGUGUUUUCCUUAACUCUUUCUGGCCAGUUGACCGUGCUAAGUUGGGCCAUAAAUCGGCAGU ...((((......))))..((((((((((((..((((((...(------(...(((..((.(((......))).))..))))).))))))..))))))))))))(((.......)))... ( -33.00) >DroEre_CAF1 118533 119 + 1 CAUAUCUUCUAAUAGAUCGCCAGCUUGGCAUACUUAACUUACGUGUACGUACCCAGUUAGUGUU-UCCUUAACUCUUUCUGGCCAGUUCGCCGUGCUAAGUUGGGCCAUAAAUCGGCAGU ...((((......))))..((((((((((((....((((((((....)))).((((..((.(((-.....))).))..))))..))))....))))))))))))(((.......)))... ( -35.60) >DroAna_CAF1 59443 114 + 1 CAUAUCUUCUAAUAGAUCGCCAACUUGGCAUACUUAACUUACG------UCCCGAGUUAGUGUUUUCCUUAACUCUUUCUGGCCAGUUGACCGUGCUAAGUUGGGCCAUAAAUCGGCAGU ...((((......))))..((((((((((((..((((((...(------((..(((((((.(....).))))))).....))).))))))..))))))))))))(((.......)))... ( -34.00) >consensus CAUAUCUUCUAAUAGAUCGCCAACUUGGCAUACUUAACUUACG______UACCCAGUUAGUGUUUUCCUUAACUCUUUCUGGCCAGUUGACCGUGCUAAGUUGGGCCAUAAAUCGGCAGU ...((((......))))..((((((((((((..((((((.............((((..((.(((......))).))..))))..))))))..))))))))))))(((.......)))... (-29.47 = -30.03 + 0.56)

| Location | 14,591,671 – 14,591,785 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 92.37 |
| Mean single sequence MFE | -31.87 |
| Consensus MFE | -28.45 |
| Energy contribution | -28.57 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.89 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.588666 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
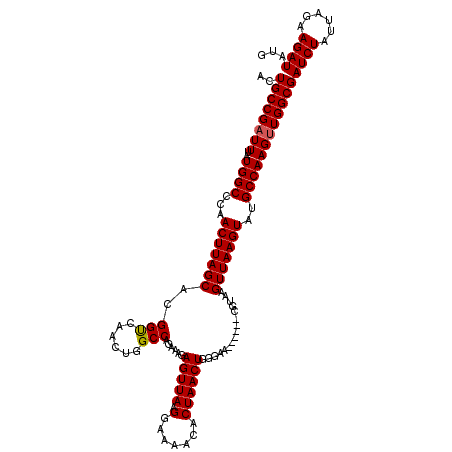
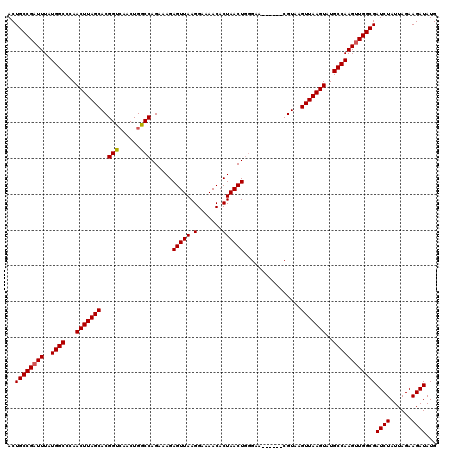
>2L_DroMel_CAF1 14591671 114 - 22407834 ACUGCCGAUUUAUGGCCCAACUUAGCACGGUCAACUGGCCAGAAAGAGUUAAGGAAAACACUAACUGCGAA------CGUAAGUUAAGUAUGCCAAGUUGGCGAUCUAUUAGAAGAUAUG ..((((((((..((((...(((((((..((((....))))......(((((.(.......)))))).....------.....)))))))..))))))))))))((((......))))... ( -30.80) >DroEre_CAF1 118533 119 - 1 ACUGCCGAUUUAUGGCCCAACUUAGCACGGCGAACUGGCCAGAAAGAGUUAAGGA-AACACUAACUGGGUACGUACACGUAAGUUAAGUAUGCCAAGCUGGCGAUCUAUUAGAAGAUAUG ..(((((.....((((...(((((((..(((......)))......(((((.(..-..)..)))))...((((....)))).)))))))..))))...)))))((((......))))... ( -30.30) >DroAna_CAF1 59443 114 - 1 ACUGCCGAUUUAUGGCCCAACUUAGCACGGUCAACUGGCCAGAAAGAGUUAAGGAAAACACUAACUCGGGA------CGUAAGUUAAGUAUGCCAAGUUGGCGAUCUAUUAGAAGAUAUG ..((((((((..((((...(((((((..((((....)))).....((((((.(.......)))))))....------.....)))))))..))))))))))))((((......))))... ( -34.50) >consensus ACUGCCGAUUUAUGGCCCAACUUAGCACGGUCAACUGGCCAGAAAGAGUUAAGGAAAACACUAACUGGGAA______CGUAAGUUAAGUAUGCCAAGUUGGCGAUCUAUUAGAAGAUAUG ..((((((((..((((...(((((((..(((......)))......(((((.(.......))))))................)))))))..))))))))))))((((......))))... (-28.45 = -28.57 + 0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:37:10 2006