
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,560,283 – 14,560,381 |
| Length | 98 |
| Max. P | 0.760867 |

| Location | 14,560,283 – 14,560,381 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 87.20 |
| Mean single sequence MFE | -44.70 |
| Consensus MFE | -34.67 |
| Energy contribution | -35.36 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.608194 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
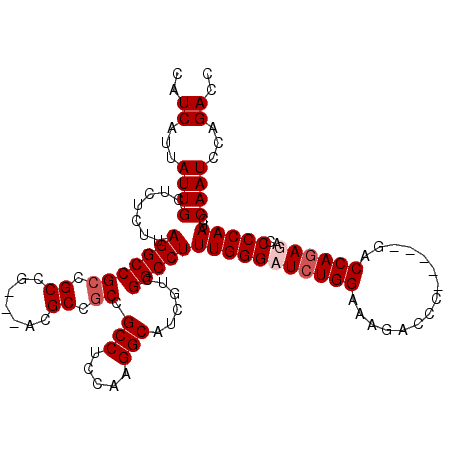
>2L_DroMel_CAF1 14560283 98 + 22407834 GGUCUGGAUUGGAUUGGGGUCUCUGGUC-----GGGUCUUUCCAGAUCCCAAAGGCCGGCGAAGCCUUGGAGGCGGC------------GGCGGCCUAAGAGAGCAAUAAUGAUG .(((...((((..(((((((((......-----(((....))))))))))))((((((.((..(((.....)))..)------------).)))))).......))))...))). ( -40.30) >DroSec_CAF1 91893 110 + 1 GGUCUGGAUUGGAUUGGGGUCUCUGGUC-----GGGUCUUUCCAGAGCCCAAAGGCCGACGAUGCCUUGGAGGCGGCGGCGUCGUCGGCGGCGGCCUAAGAGAGCAAUAAUGAUG (((((.....)))))..(.(((((((((-----(((.(((....))))))....(((((((((((((((....))).))))))))))))...))))..))))).).......... ( -47.10) >DroSim_CAF1 79359 110 + 1 GGUCUGGAUUGGAUUGGGGUCUCUGGUC-----GGGUCUUUCCAGAGCCCAAAGGCCGACGAUGCCUUGGAGGCGGCGGCGUCGUCGGCGGCGGCCUAAGAGAGCAAUAAUGAUG (((((.....)))))..(.(((((((((-----(((.(((....))))))....(((((((((((((((....))).))))))))))))...))))..))))).).......... ( -47.10) >DroYak_CAF1 83330 112 + 1 GGUCUGGAUUGGAUUGGGCUCUCUGGUCGGGUCGUGUCUUUCCACAUCCCAAAGGCCCACGUUGCCUUGGAGGCGGUGGCGG---UGGCGGCGGCCUAAGAGAGCAAUAAUGAUG (((((.(((..((........))..)))(((..(((......)))..)))..)))))...((((((((..((((.((.((..---..)).)).))))..))).)))))....... ( -44.30) >consensus GGUCUGGAUUGGAUUGGGGUCUCUGGUC_____GGGUCUUUCCAGAGCCCAAAGGCCGACGAUGCCUUGGAGGCGGCGGCGU___CGGCGGCGGCCUAAGAGAGCAAUAAUGAUG (((((........(((((..((((((...............))))))))))))))))......(((((..((((.((.((.......)).)).))))..))).)).......... (-34.67 = -35.36 + 0.69)



| Location | 14,560,283 – 14,560,381 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 87.20 |
| Mean single sequence MFE | -35.38 |
| Consensus MFE | -24.03 |
| Energy contribution | -25.16 |
| Covariance contribution | 1.13 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.12 |
| Structure conservation index | 0.68 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.760867 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14560283 98 - 22407834 CAUCAUUAUUGCUCUCUUAGGCCGCC------------GCCGCCUCCAAGGCUUCGCCGGCCUUUGGGAUCUGGAAAGACCC-----GACCAGAGACCCCAAUCCAAUCCAGACC .......((((.(((((.((((((.(------------(..(((.....)))..)).))))))(((((.(((....))))))-----))..)))))...))))............ ( -32.40) >DroSec_CAF1 91893 110 - 1 CAUCAUUAUUGCUCUCUUAGGCCGCCGCCGACGACGCCGCCGCCUCCAAGGCAUCGUCGGCCUUUGGGCUCUGGAAAGACCC-----GACCAGAGACCCCAAUCCAAUCCAGACC .......((((.(((((..((.....((((((((.(((...........))).))))))))..(((((.(((....))))))-----)))))))))...))))............ ( -36.30) >DroSim_CAF1 79359 110 - 1 CAUCAUUAUUGCUCUCUUAGGCCGCCGCCGACGACGCCGCCGCCUCCAAGGCAUCGUCGGCCUUUGGGCUCUGGAAAGACCC-----GACCAGAGACCCCAAUCCAAUCCAGACC .......((((.(((((..((.....((((((((.(((...........))).))))))))..(((((.(((....))))))-----)))))))))...))))............ ( -36.30) >DroYak_CAF1 83330 112 - 1 CAUCAUUAUUGCUCUCUUAGGCCGCCGCCA---CCGCCACCGCCUCCAAGGCAACGUGGGCCUUUGGGAUGUGGAAAGACACGACCCGACCAGAGAGCCCAAUCCAAUCCAGACC ..........(((((((.(((((...((..---..))(((.(((.....)))...))))))))(((((.((((......)))).)))))..)))))))................. ( -36.50) >consensus CAUCAUUAUUGCUCUCUUAGGCCGCCGCCG___ACGCCGCCGCCUCCAAGGCAUCGUCGGCCUUUGGGAUCUGGAAAGACCC_____GACCAGAGACCCCAAUCCAAUCCAGACC ..((...((((.......(((((((.((.......)).)).(((.....)))......)))))(((((((((((...............))))))..)))))..))))...)).. (-24.03 = -25.16 + 1.13)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:36:39 2006