
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,408,157 – 14,408,316 |
| Length | 159 |
| Max. P | 0.971144 |

| Location | 14,408,157 – 14,408,276 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 80.39 |
| Mean single sequence MFE | -29.16 |
| Consensus MFE | -21.56 |
| Energy contribution | -23.50 |
| Covariance contribution | 1.94 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.26 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.67 |
| SVM RNA-class probability | 0.971144 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
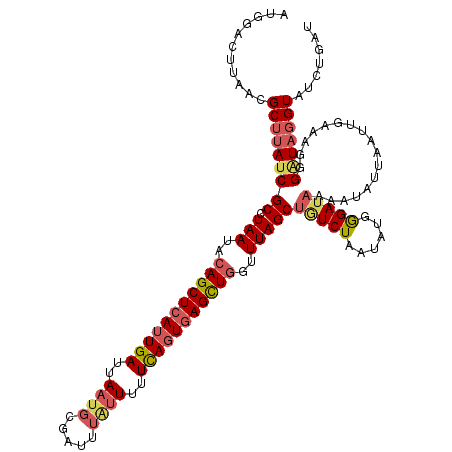
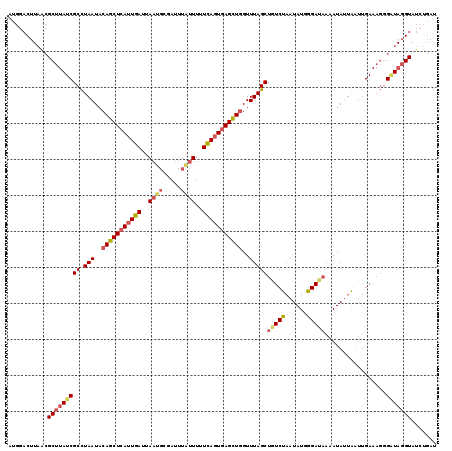
>2L_DroMel_CAF1 14408157 119 - 22407834 AUGGACUUAGCGCUUAUCGCCUAAUACAGCUCAGUGAUUAUUGCGAUUUCUUUUUCAGUGAGUUGGUUUAGCUAUCUAAUAUGGGAUAAAAUAUUAAUUGAAAGGGAUAGGUAUCUGAU ......((((.(((((((.(((....(((((((.(((.................))).))))))).......(((((......)))))..............))))))))))..)))). ( -27.43) >DroSec_CAF1 29235 119 - 1 AUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAUAUUGGAUAAAAUAUGAAUAGAAAGGGUUAGGUAUCUGAU ....(((((((.(((.((((.(((..(((((((((((..((((.....))))..)))))))))))..)))))((((((....))))))...........))))).)))))))....... ( -36.50) >DroSim_CAF1 28093 119 - 1 AUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAUAUAAGAUAAAAUAUUGGUUGAAAGGGAUAGGUAUCUGAU ...........(((((((.(((....(((((((((((..((((.....))))..))))))))))).(((((((((((......))))........)))))))))))))))))....... ( -34.80) >DroEre_CAF1 29424 100 - 1 -------------UUAUCGCUUAAUACAGCUCAUUGAUUAAUGCGAUUGGUUUUUUAGUAAGCUUGUUUAGCUGUCUAAUAUGGGAAUUAAUAAUAAUUGAA-UUGGU-----UCUGCG -------------..((((((((((.(((....)))))))).))))).......((((..((((.....))))..))))....(((((((((.........)-)))))-----)))... ( -17.90) >consensus AUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAUAUGGGAUAAAAUAUUAAUUGAAAGGGAUAGGUAUCUGAU ...........(((((((((.(((..(((((((((((..((((.....))))..)))))))))))..)))))(((((......))))).................)))))))....... (-21.56 = -23.50 + 1.94)

| Location | 14,408,196 – 14,408,316 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.11 |
| Mean single sequence MFE | -27.71 |
| Consensus MFE | -25.47 |
| Energy contribution | -26.03 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.92 |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.619795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
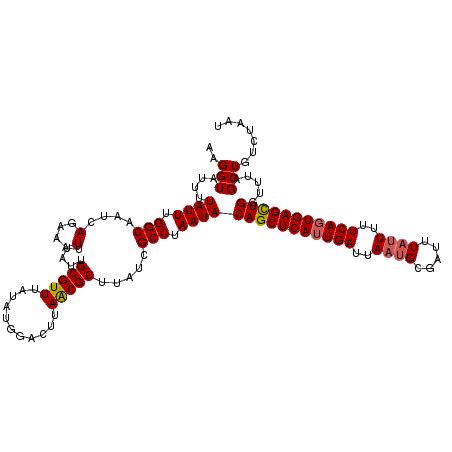
>2L_DroMel_CAF1 14408196 120 - 22407834 AAGGUAUUUUAUUCGGCAAUCAGAAAUUUAUUGCGUUUAUAUGGACUUAGCGCUUAUCGCCUAAUACAGCUCAGUGAUUAUUGCGAUUUCUUUUUCAGUGAGUUGGUUUAGCUAUCUAAU .(((((.........(((((.(....)..))))).......(((((((((.((.....))))))..(((((((.(((.................))).))))))))))))..)))))... ( -23.33) >DroSec_CAF1 29274 120 - 1 AAGGUAUUUUAUUUGGCAAUCAGAAAUUUAUUGCGUUUAUAUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAU ...............(((((.(....)..))))).......(((((.....((.....))((((..(((((((((((..((((.....))))..)))))))))))..))))..))))).. ( -29.90) >DroSim_CAF1 28132 120 - 1 AAGGUAUUUUAUUUGGCAAUCAGAAAUUUAUUGCGUUUAUAUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAU ...............(((((.(....)..))))).......(((((.....((.....))((((..(((((((((((..((((.....))))..)))))))))))..))))..))))).. ( -29.90) >consensus AAGGUAUUUUAUUUGGCAAUCAGAAAUUUAUUGCGUUUAUAUGGACUUAACGCUUAUCGCCUAAUACAGCUCAUUGAUUAAUGCGAUUUAUUUUUCAGUGAGCUGGUUUAGCUGUCUAAU ..(((....((((.(((....(....).....(((((...........))))).....))).))))(((((((((((..((((.....))))..))))))))))).....)))....... (-25.47 = -26.03 + 0.56)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:33:41 2006