| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 212,648 – 212,749 |
| Length | 101 |
| Max. P | 0.616941 |

| Location | 212,648 – 212,749 |
|---|---|
| Length | 101 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 79.81 |
| Mean single sequence MFE | -32.99 |
| Consensus MFE | -18.28 |
| Energy contribution | -19.06 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.91 |
| Structure conservation index | 0.55 |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.616941 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 212648 101 - 22407834 AUCGGGGUCCUCUUCCAGUUCAGCAGCUGCUGCGGGACUGGCAGCUAAUCUGCCCAGUCAAACUACAAUAACCAUAGCUCCUCCCAUUAGCAGUGGAGCCA ...((((((((.(..((((......))))..).))))))(((((.....))))).................))...(((((.............))))).. ( -28.42) >DroPse_CAF1 6250 98 - 1 AGGAUCCUCCUCAUCGAACGCGGCAGCUGCU---GGCCUGGCGGCCAAUCUGCCGUCGCAGACUACCAUCACCAUAGCGCCGCCCAUCAGCAGUGGAGCCA (((.....)))..........(((.((((((---((...((((((...(((((....)))))(((.........))).))))))..))))))))...))). ( -36.60) >DroSim_CAF1 4925 101 - 1 AUCAGGCUCCUCUUCCAGUUCAGCAGCUGCUGCGGGACUGGCAGCUAAUCUGCCCAGUCAAACUACAAUAACCAUAGCUCCUCCCAUUAGCAGUGGAGCCA ....((((((.......(.....).(((((((.(((((((((((.....))).))))))...(((.........)))......))..))))))))))))). ( -33.00) >DroWil_CAF1 6503 98 - 1 UGGCGGCAAUUCUUCCUCCUCGGCAGCUGCU---GGCAUAGCCGCCAAUCUGCCAUCGCAGACCACCAUCACAAUUGCUCCGCCCAUUAGCAGUGGAGCCA (((((((.......((.....))..((....---.))...))))))).(((((....)))))..............(((((((.........))))))).. ( -32.70) >DroYak_CAF1 6985 101 - 1 GUCAGGCUCUUCUUCCAGUUCAGCAGCUGCUGCGGGACUGGCAGCUAAUCUGCCCAGUCAAACUACAAUAACCAUAGCUCCUCCCAUUAGCAGUGGAGCCA ....(((((.......((((((((....))))...(((((((((.....))).)))))).))))........(((.(((.........))).)))))))). ( -30.60) >DroPer_CAF1 6379 98 - 1 AGGAUCCUCCUCAUCGAACGCGGCAGCUGCU---GGCCUGGCGGCCAAUCUGCCGUCGCAGACUACCAUCACCAUAGCGCCGCCCAUCAGCAGUGGAGCCA (((.....)))..........(((.((((((---((...((((((...(((((....)))))(((.........))).))))))..))))))))...))). ( -36.60) >consensus AGCAGGCUCCUCUUCCAGCUCAGCAGCUGCU___GGACUGGCAGCCAAUCUGCCCACGCAAACUACAAUAACCAUAGCUCCGCCCAUUAGCAGUGGAGCCA ....((((((...............((((((...(((..(((((.....)))))........(((.........)))....)))....)))))))))))). (-18.28 = -19.06 + 0.78)

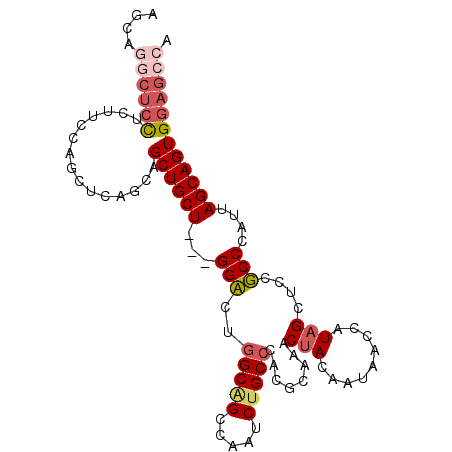
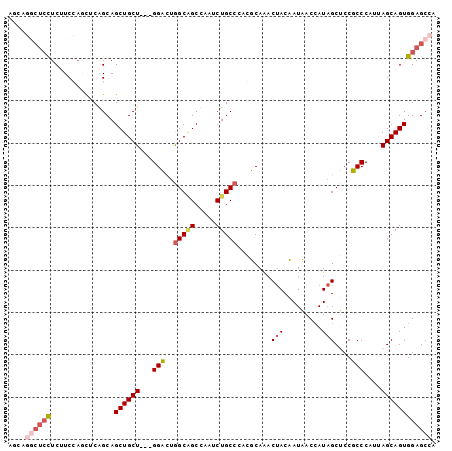
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:24:11 2006