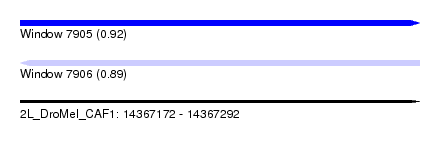

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,367,172 – 14,367,292 |
| Length | 120 |
| Max. P | 0.922012 |

| Location | 14,367,172 – 14,367,292 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 87.20 |
| Mean single sequence MFE | -34.78 |
| Consensus MFE | -32.46 |
| Energy contribution | -32.90 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.14 |
| SVM RNA-class probability | 0.922012 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
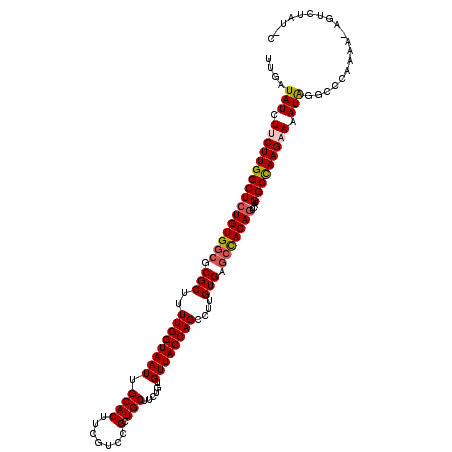
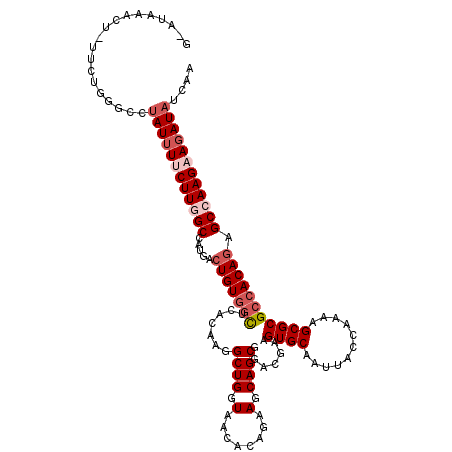
>2L_DroMel_CAF1 14367172 120 + 22407834 UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCGUCCGCUGCUUCUGUGUUACCAGCCUUGUGAGCCACAGUCAUGGCUAAGAAAAUAGGCCCACUGAUGUGUAUGC ....(((.((((((((((((((((.(((..((((((((.((((.......).)))......))))))))....))).)))))))....))))))))).)))....(((....)))..... ( -37.10) >DroSec_CAF1 15142 120 + 1 UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCGUCCGCUGCUUCUGUGUUACCAGCCUCGUGAGCCACAGUCAUGGCUAAGAAAAUAGGCCCUGAAUAGUUUAUAC ........((((((((((((((((.(((..((((((((.((((.......).)))......))))))))....))).)))))))....))))))))).((((((........)))))).. ( -35.80) >DroSim_CAF1 17253 120 + 1 UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCGUCCGCUGCUUCUGUGUUACCAGCCUCGUGAGCCACAGUCAGGGCCAAGAAAAUGGGCCCUGAAUAGUUUAUAC .............(((((((((((.(((..((((((((.((((.......).)))......))))))))....))).)))))))((((((((.........))))))))..))))..... ( -38.90) >DroEre_CAF1 16157 115 + 1 UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCAUCCGCUGCUUCUGUGUUACCAGCAUUGUGAGCCACAGUCAUGGCCAAGCAUAUAAUAUCAAAA-AAUCU---- (((((((...((((((((((((((.(((..((((((((.((((.......).)))......))))))))....))).)))))))....)))))))......)))))))..-.....---- ( -37.80) >DroYak_CAF1 15371 115 + 1 UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCGUCCGCUGCUUCUGUGUUACCAGCCUUGUGAAGUACACUCAUGGCCAAGCAAAUAAAAUCAAAA-AAUCU---- (((((.....(((((((..(((.((((...((((((((.((((.......).)))......))))))))...)))).....)))....)))))))........)))))..-.....---- ( -24.32) >consensus UUGAUAUCUUCUUGGCUCUGUGGCGCGCUUUUGGUAAUUGCACUUCGUCCGCUGCUUCUGUGUUACCAGCCUUGUGAGCCACAGUCAUGGCCAAGAAAAUAGGCCCAAAA_AGUCUAU_C ....(((.((((((((((((((((.(((..((((((((.((((.......).)))......))))))))....))).)))))))....))))))))).)))................... (-32.46 = -32.90 + 0.44)



| Location | 14,367,172 – 14,367,292 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.20 |
| Mean single sequence MFE | -33.62 |
| Consensus MFE | -28.46 |
| Energy contribution | -30.10 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.893698 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14367172 120 - 22407834 GCAUACACAUCAGUGGGCCUAUUUUCUUAGCCAUGACUGUGGCUCACAAGGCUGGUAACACAGAAGCAGCGGACGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ((...(((....))).)).(((((((((.((.....(((((((.......((((.(........).)))).......((((...........))))))))))).)).))))))))).... ( -34.40) >DroSec_CAF1 15142 120 - 1 GUAUAAACUAUUCAGGGCCUAUUUUCUUAGCCAUGACUGUGGCUCACGAGGCUGGUAACACAGAAGCAGCGGACGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ...................(((((((((.((.....(((((((((.((..(((.(.....)...)))..))))....((((...........))))))))))).)).))))))))).... ( -31.00) >DroSim_CAF1 17253 120 - 1 GUAUAAACUAUUCAGGGCCCAUUUUCUUGGCCCUGACUGUGGCUCACGAGGCUGGUAACACAGAAGCAGCGGACGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ...........((((((((.........))))))))(((((((((.((..(((.(.....)...)))..))))....((((...........)))))))))))................. ( -38.00) >DroEre_CAF1 16157 115 - 1 ----AGAUU-UUUUGAUAUUAUAUGCUUGGCCAUGACUGUGGCUCACAAUGCUGGUAACACAGAAGCAGCGGAUGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ----.....-..((((((((.....((((((.....((((((((((...(((((.(........).)))))..))).((((...........))))))))))).))))))..)))))))) ( -37.80) >DroYak_CAF1 15371 115 - 1 ----AGAUU-UUUUGAUUUUAUUUGCUUGGCCAUGAGUGUACUUCACAAGGCUGGUAACACAGAAGCAGCGGACGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ----.....-..((((...(((((.((((((..((.(((.....)))...((((.(........).))))((.....((((...........)))).)).))..)))))).))))))))) ( -26.90) >consensus G_AUAAACU_UUCUGGGCCUAUUUUCUUGGCCAUGACUGUGGCUCACAAGGCUGGUAACACAGAAGCAGCGGACGAAGUGCAAUUACCAAAAGCGCGCCACAGAGCCAAGAAGAUAUCAA ...................((((((((((((.....(((((((.......((((.(........).)))).......((((...........))))))))))).)))))))))))).... (-28.46 = -30.10 + 1.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:38 2006