
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,366,981 – 14,367,133 |
| Length | 152 |
| Max. P | 0.899901 |

| Location | 14,366,981 – 14,367,101 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.92 |
| Mean single sequence MFE | -32.02 |
| Consensus MFE | -28.42 |
| Energy contribution | -28.70 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.89 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.899901 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
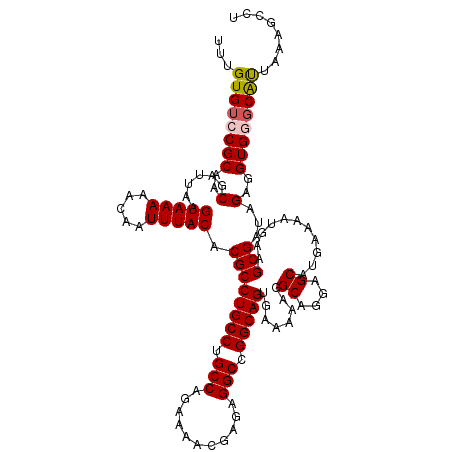
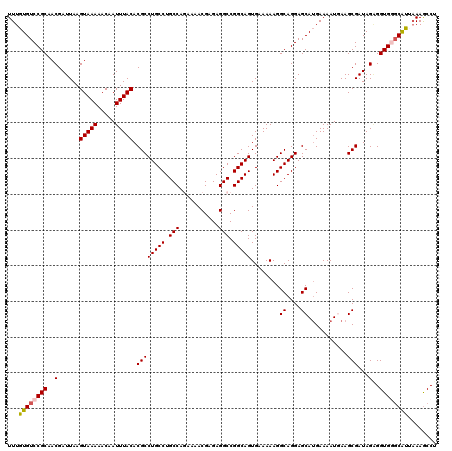
>2L_DroMel_CAF1 14366981 120 - 22407834 UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCAGAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAUGAAAAUGAAGCGAUAGUGGUGGCCAUUAAAGCCU ((..(((((...........(((((.....)))))..((((.((((((.(....(....).).))))).)....)))).))).))..))......((..((((((...))))))..)).. ( -32.00) >DroSec_CAF1 14957 120 - 1 UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCAGAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAUGAAAAUCAAGCGAUAGAGGUGGGCACUAAAGCCU ...((((((((....(((..(((((.....)))))...((((((((((.(....(....).).)))..(....)))))))).((.((.....)).)))))....))))))))........ ( -35.30) >DroSim_CAF1 17068 120 - 1 UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCAGAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAUGAAAAUCAAGCGAUAGAGGUGGGCAUUCAAGCCU ...((((((((....(((..(((((.....)))))...((((((((((.(....(....).).)))..(....)))))))).((.((.....)).)))))....))))))))........ ( -33.10) >DroEre_CAF1 15972 120 - 1 UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCACAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAAGAAAAUGAAGCGAUAGGGGUGAGCGUGAAAACCC ((..((..(((..(......(((((.....))))).((((((((.(((.(.......).))).)))))........((....))...........)))...)..)))..))..))..... ( -29.90) >DroYak_CAF1 15177 120 - 1 UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCACAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAUGAAAAUGAAGCGAUAGAGGUGAGCAUGAAAGCCC ((..((..(((..(......(((((.....))))).((((((((.(((.(.......).))).)))))...............(((....)))..)))...)..)))..))..))..... ( -29.80) >consensus UUUGUGUCCGCAACGAUUAAGUAAAAACAAUUUACACGCCUGCCUGCCAGAAAACGAGAGGCCGGCAGUGAAAAAGGCAGGAGCAUGAAAAUGAAGCGAUAGAGGUGGGCAUUAAAGCCU ...((((((((..(......(((((.....))))).((((((((.(((...........))).)))))........((....))...........)))...)..))))))))........ (-28.42 = -28.70 + 0.28)

| Location | 14,367,021 – 14,367,133 |
|---|---|
| Length | 112 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.59 |
| Mean single sequence MFE | -30.94 |
| Consensus MFE | -29.20 |
| Energy contribution | -29.20 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.02 |
| SVM RNA-class probability | 0.545230 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
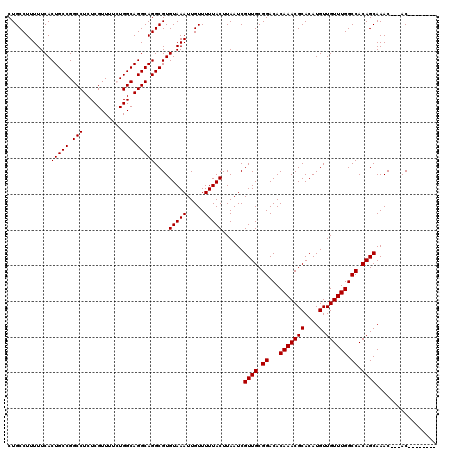
>2L_DroMel_CAF1 14367021 112 + 22407834 CUGCCUUUUUCACUGCCGGCCUCUCGUUUUCUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAACAGCAC-------- ............(((((.(((...........))).)))))((..(((((.....)))))......((((.((...(((((((....).)))))).)).)))).....))..-------- ( -31.60) >DroSec_CAF1 14997 109 + 1 CUGCCUUUUUCACUGCCGGCCUCUCGUUUUCUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAAC---AC-------- ..((((......(((((((...........)))))))..))))(((((((.....)))))......((((.((...(((((((....).)))))).)).))))..))---..-------- ( -29.80) >DroSim_CAF1 17108 109 + 1 CUGCCUUUUUCACUGCCGGCCUCUCGUUUUCUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAAC---AC-------- ..((((......(((((((...........)))))))..))))(((((((.....)))))......((((.((...(((((((....).)))))).)).))))..))---..-------- ( -29.80) >DroEre_CAF1 16012 109 + 1 CUGCCUUUUUCACUGCCGGCCUCUCGUUUUGUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAAC---AC-------- ............(((((.(((.(.......).))).)))))..(((((((.....)))))......((((.((...(((((((....).)))))).)).))))..))---..-------- ( -30.30) >DroYak_CAF1 15217 117 + 1 CUGCCUUUUUCACUGCCGGCCUCUCGUUUUGUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAAC---ACAGCACACA .(((........(((((.(((.(.......).))).)))))..((((.....((....))......((((.((...(((((((....).)))))).)).))))..))---)).))).... ( -33.20) >consensus CUGCCUUUUUCACUGCCGGCCUCUCGUUUUCUGGCAGGCAGGCGUGUAAAUUGUUUUUACUUAAUCGUUGCGGACACAAACGCACAUGUUGUUUGGCCACAGCAAAC___AC________ ............(((((.(((...........))).)))))....(((((.....)))))......((((.((...(((((((....).)))))).)).))))................. (-29.20 = -29.20 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:36 2006