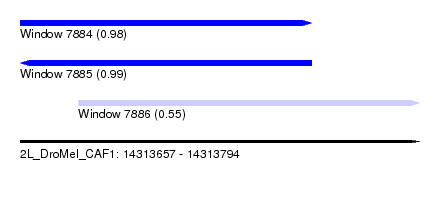
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,313,657 – 14,313,794 |
| Length | 137 |
| Max. P | 0.987505 |

| Location | 14,313,657 – 14,313,757 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 72.48 |
| Mean single sequence MFE | -17.19 |
| Consensus MFE | -12.66 |
| Energy contribution | -12.42 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.74 |
| SVM decision value | 1.94 |
| SVM RNA-class probability | 0.983451 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14313657 100 + 22407834 CCGGCCAA--------------------ACGGCAUCAACUUCAACACCAACACCAAAACCGGCAUUAAACUGAGGCCAGUUACUGUUACAUGACACACGGUGUCAAAAUAUGAACAGUAA ..((((..--------------------..((..............))....((......))...........))))..((((((((.((((((((...)))))).....)))))))))) ( -21.54) >DroPse_CAF1 180269 114 + 1 ACAUCCAAUAGCAACAGCAACAGCAACAUCAG------CAUCAACACCAACACCGAAACCGACCAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAA ..........((..(((.....((.......)------)..............((....))........)))..))...((((((((.((((((((...)))))).....)))))))))) ( -16.30) >DroWil_CAF1 218336 88 + 1 ------AU--------------------AAGG---CUACAUCAACACCAACACCAUAUCC---AUAAAGCUAAAGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGCAA ------..--------------------..((---((.......................---..........)))).....(((((.((((((((...)))))).....)))))))... ( -11.01) >DroYak_CAF1 137573 100 + 1 CCAGCCAA--------------------ACGGCAUCAACUUCAACACCAACACCAAAACCGGCAUUAAACUGAGGCCAGUUACUGUUACAUGACACACGGUGUCAAAAUAUGAACAGUAA ...(((..--------------------..)))...........................(((.((.....)).)))..((((((((.((((((((...)))))).....)))))))))) ( -20.70) >DroAna_CAF1 150385 85 + 1 CCGGAGG-----------------------------------AACACCAACACCAAAACCGGCAAUAAACUGAGCCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAA ((((.((-----------------------------------..........))....)))).....(((((....)))))((((((.((((((((...)))))).....)))))))).. ( -17.00) >DroPer_CAF1 185718 120 + 1 ACAUCCAAUAGCAACAGCAACAGCAACAUCAGCAUCAGCAUCAACACCAACACCGAAACCGACCAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAA ................((..(((........((....))..............((....))........)))..))...((((((((.((((((((...)))))).....)))))))))) ( -16.60) >consensus CCAGCCAA____________________ACGG___CAACAUCAACACCAACACCAAAACCGGCAAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAA ...................................................................(((((....)))))((((((.((((((((...)))))).....)))))))).. (-12.66 = -12.42 + -0.25)


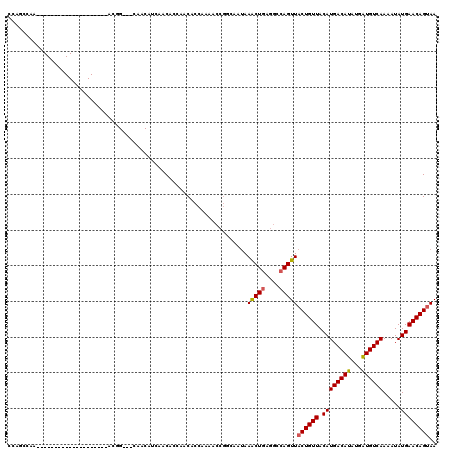
| Location | 14,313,657 – 14,313,757 |
|---|---|
| Length | 100 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 72.48 |
| Mean single sequence MFE | -29.17 |
| Consensus MFE | -15.36 |
| Energy contribution | -15.30 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.53 |
| SVM decision value | 2.08 |
| SVM RNA-class probability | 0.987505 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
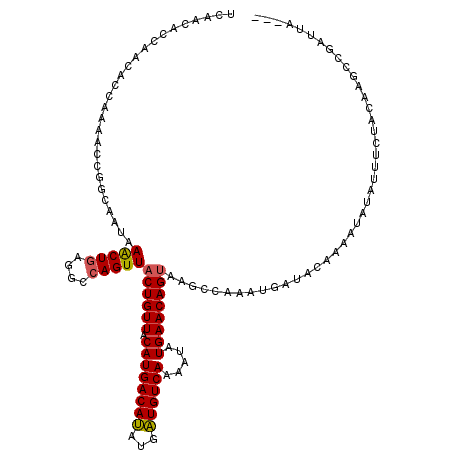
>2L_DroMel_CAF1 14313657 100 - 22407834 UUACUGUUCAUAUUUUGACACCGUGUGUCAUGUAACAGUAACUGGCCUCAGUUUAAUGCCGGUUUUGGUGUUGGUGUUGAAGUUGAUGCCGU--------------------UUGGCCGG ((((((((((.....((((((...)))))))).))))))))((((((.......(((((((....)))))))((((((......))))))..--------------------..)))))) ( -32.50) >DroPse_CAF1 180269 114 - 1 UUACUGUUCAUAUUUUGACAUCAUAUGUCAUGUAACAGUAACUGGCCUCAGUUUAUGGUCGGUUUCGGUGUUGGUGUUGAUG------CUGAUGUUGCUGUUGCUGUUGCUAUUGGAUGU .....(((((.....((((((...)))))).(((((((((((.((((.........))))(((.((((((((......))))------))))....))))))))))))))...))))).. ( -34.10) >DroWil_CAF1 218336 88 - 1 UUGCUGUUCAUAUUUUGACAUCAUAUGUCAUGUAACAGUAACUGGCUUUAGCUUUAU---GGAUAUGGUGUUGGUGUUGAUGUAG---CCUU--------------------AU------ ..(((..(((....(..(((((((((.(((((...(((...)))((....))..)))---)))))))))))..)...)))...))---)...--------------------..------ ( -20.70) >DroYak_CAF1 137573 100 - 1 UUACUGUUCAUAUUUUGACACCGUGUGUCAUGUAACAGUAACUGGCCUCAGUUUAAUGCCGGUUUUGGUGUUGGUGUUGAAGUUGAUGCCGU--------------------UUGGCUGG ((((((((((.....((((((...)))))))).))))))))..(((.(((((((((((((..(......)..))))))))).)))).)))..--------------------........ ( -30.70) >DroAna_CAF1 150385 85 - 1 UUACUGUUCAUAUUUUGACAUCAUAUGUCAUGUAACAGUAACUGGGCUCAGUUUAUUGCCGGUUUUGGUGUUGGUGUU-----------------------------------CCUCCGG .(((((((((.....((((((...)))))))).)))))))((((....))))......((((....((..........-----------------------------------)).)))) ( -18.20) >DroPer_CAF1 185718 120 - 1 UUACUGUUCAUAUUUUGACAUCAUAUGUCAUGUAACAGUAACUGGCCUCAGUUUAUGGUCGGUUUCGGUGUUGGUGUUGAUGCUGAUGCUGAUGUUGCUGUUGCUGUUGCUAUUGGAUGU .....(((((.....((((((...)))))).(((((((((((.((((.........))))(((.(((((((..((......))..)))))))....))))))))))))))...))))).. ( -38.80) >consensus UUACUGUUCAUAUUUUGACAUCAUAUGUCAUGUAACAGUAACUGGCCUCAGUUUAAUGCCGGUUUUGGUGUUGGUGUUGAUGUUG___CCGU____________________UUGGCUGG .(((((((((.....((((((...)))))))).)))))))((..((.((((.............)))).))..))............................................. (-15.36 = -15.30 + -0.05)



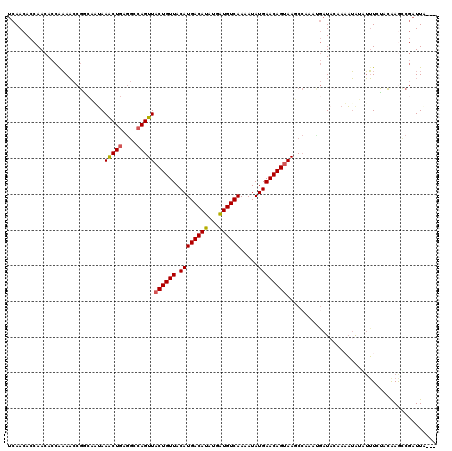
| Location | 14,313,677 – 14,313,794 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.73 |
| Mean single sequence MFE | -20.13 |
| Consensus MFE | -12.66 |
| Energy contribution | -12.42 |
| Covariance contribution | -0.25 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.03 |
| SVM RNA-class probability | 0.549595 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14313677 117 + 22407834 UCAACACCAACACCAAAACCGGCAUUAAACUGAGGCCAGUUACUGUUACAUGACACACGGUGUCAAAAUAUGAACAGUAAUCCAAACGAUACAAAAUAUAUUUGCACAAACCGAUUA--- ...................(((...........((...(((((((((.((((((((...)))))).....)))))))))))))........((((.....))))......)))....--- ( -18.50) >DroPse_CAF1 180303 117 + 1 UCAACACCAACACCGAAACCGACCAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAAGCCAAAUGAUACAAUAUGGGUUUCUAUGAGCCGAUUA--- ..........((..((((((...((.....)).(((...((((((((.((((((((...)))))).....)))))))))))))...............))))))..)).........--- ( -22.90) >DroWil_CAF1 218347 112 + 1 UCAACACCAACACCAUAUCC---AUAAAGCUAAAGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGCAAUCGG-AUGAUGUAAAUGAUAAAU----GAGUGGAUUUUCU ..........((((((...(---((...((.....((.(((.(((((.((((((((...)))))).....))))))).))).))-.....))..)))....))----).)))........ ( -16.80) >DroYak_CAF1 137593 117 + 1 UCAACACCAACACCAAAACCGGCAUUAAACUGAGGCCAGUUACUGUUACAUGACACACGGUGUCAAAAUAUGAACAGUAAUCCAAACGAUACAAAAUAUAUUUGCACAAACCGAUUU--- ...................(((...........((...(((((((((.((((((((...)))))).....)))))))))))))........((((.....))))......)))....--- ( -18.50) >DroAna_CAF1 150392 112 + 1 --AACACCAACACCAAAACCGGCAAUAAACUGAGCCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAAGCCAAAAAAUU--AAAUG-GUUUUUACAAGUCCAUUC--- --..................(((....(((((....)))))((((((.((((((((...)))))).....))))))))..)))........--.((((-(...........))))).--- ( -21.20) >DroPer_CAF1 185758 117 + 1 UCAACACCAACACCGAAACCGACCAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAAGCCAAAUGAUACAAUAUGGGUUUCUAUGAGCCGAUUA--- ..........((..((((((...((.....)).(((...((((((((.((((((((...)))))).....)))))))))))))...............))))))..)).........--- ( -22.90) >consensus UCAACACCAACACCAAAACCGGCAAUAAACUGAGGCCAGUUACUGUUACAUGACAUAUGAUGUCAAAAUAUGAACAGUAAGCCAAAUGAUACAAAAUAUAUUUCUACAAGCCGAUUA___ ...........................(((((....)))))((((((.((((((((...)))))).....)))))))).......................................... (-12.66 = -12.42 + -0.25)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:32:18 2006