
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 14,069,184 – 14,069,334 |
| Length | 150 |
| Max. P | 0.802403 |

| Location | 14,069,184 – 14,069,282 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 78.14 |
| Mean single sequence MFE | -25.52 |
| Consensus MFE | -11.45 |
| Energy contribution | -14.15 |
| Covariance contribution | 2.70 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.802403 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14069184 98 - 22407834 AGCAGGAAAAAAACCGGCGAAACCGAAAGUUCCUUGCUGACACAAGGCUGGCGGUAAUUAAAACAGGAAAUUAAAGGAAUGCUU-UUAAUUUCGUGUUC .((.((.......)).))...((((..((..(((((......)))))))..))))......((((.(((((((((((....)))-)))))))).)))). ( -29.30) >DroSec_CAF1 157947 97 - 1 AGCAGGAAA-AAACCGGUGAAACCGAAAGUUCCUUGCUGACACAAGGCUGGCGGUAAUUAAAACAGGAAAUUAAAGGAAUGCUU-UUAAUUUCGUGUUC ((((((((.-....(((.....)))....)))).))))(((((....(((.............)))(((((((((((....)))-))))))))))))). ( -27.92) >DroSim_CAF1 158880 97 - 1 AGCAGGAAA-AAACCGGUGAAACCGAAAGUUCCUUGCUGACACAAGGCUGGCGGUAAUUAAAACAGGAAAUUAAAGGAAUGCUU-UUAAUUUCGUGUUC ((((((((.-....(((.....)))....)))).))))(((((....(((.............)))(((((((((((....)))-))))))))))))). ( -27.92) >DroEre_CAF1 160796 96 - 1 AGCUG-AAA-AAACUGGCGAAACCGAAAGUUCCUUGCUGACACAAGGCUGGCGGUAAUUAAAACAGGAAAUUAAAGGAGUGCUU-UUAAUUUCGUGUUC .(((.-...-.....)))...((((..((..(((((......)))))))..))))......((((.(((((((((((....)))-)))))))).)))). ( -27.60) >DroYak_CAF1 168289 96 - 1 GGCUA-AAA-AAACUGGUGAAACCGAAAGUUUCUUGCUGACACAACUCUGGCGGUAAUUAAAACAGGACCUUAAAGGAGUGCUU-UUAAUUUCGUGUUC .((((-...-....))))((((.(....)))))((((((.((......)).))))))....((((.((..(((((((....)))-))))..)).)))). ( -17.50) >DroAna_CAF1 155492 78 - 1 -----GUGG-AAGCUUGUGAAACUGAAAGUUUUCUUUUCACAUGAGGC---------------CCGGAAAUUUAAGGGAUGCCCACUAACGUUUCCACC -----((((-((((.(((((((..(((....))).)))))))((.(((---------------((..........))...))))).....)))))))). ( -22.90) >consensus AGCAGGAAA_AAACCGGUGAAACCGAAAGUUCCUUGCUGACACAAGGCUGGCGGUAAUUAAAACAGGAAAUUAAAGGAAUGCUU_UUAAUUUCGUGUUC .....................((((..((..(((((......)))))))..))))......((((.(((((((((((....))).)))))))).)))). (-11.45 = -14.15 + 2.70)


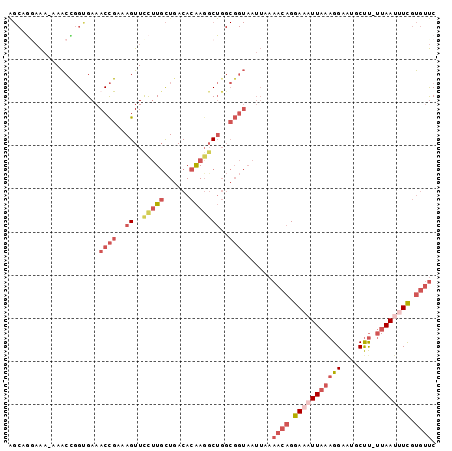
| Location | 14,069,244 – 14,069,334 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 82.19 |
| Mean single sequence MFE | -23.52 |
| Consensus MFE | -16.86 |
| Energy contribution | -17.45 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.13 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.800014 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 14069244 90 - 22407834 CAGGGAUUCUUUUUUUCACCUGCCUACUUUAAGGUGCCCAUUCAAAAGGAAAAGCAGGAAAAAAACCGGCGAAACCGAAAGUUCCUUGCU ((((((..((((......(((((.((((....)))).((........))....)))))........(((.....))))))).)))))).. ( -20.70) >DroSec_CAF1 158007 89 - 1 CAGGGAUUCUUUUUUUCACCUGCCUACUUUAAGGCACCCAUUCAAAAGGAAAAGCAGGAAA-AAACCGGUGAAACCGAAAGUUCCUUGCU ((((((..(((((((((((((((((......))))).((........)).......((...-...)))))))))..))))).)))))).. ( -26.20) >DroSim_CAF1 158940 89 - 1 CAGGGAUUCUUUUUUUCACCUGCCUACUUUAAGGCGCCCAUUCAAAAGGAAAAGCAGGAAA-AAACCGGUGAAACCGAAAGUUCCUUGCU ((((((..((((((((((((.((((......))))((...(((.....)))..)).((...-...)))))))))..))))).)))))).. ( -26.60) >DroEre_CAF1 160856 87 - 1 CAGGGAUUCUUU-UUUCACCUGCCUACUUUAAGGCGCCUGUUCAAAAGGAUGAGCUG-AAA-AAACUGGCGAAACCGAAAGUUCCUUGCU ((((((..((((-((((.((.((((......))))((..((((....))))..))..-...-.....)).)))...))))).)))))).. ( -23.70) >DroYak_CAF1 168349 87 - 1 CAGGGAUUCUUU-UUCCACCCGCCUACUUUAAGGCACCUAUUCAAAAGGAUGGGCUA-AAA-AAACUGGUGAAACCGAAAGUUUCUUGCU ((((((..((((-(..((((.((((......)))).(((((((....)))))))...-...-.....)))).....))))).)))))).. ( -23.60) >DroAna_CAF1 155538 81 - 1 CAGGUAUUCUUU-UUUCAGCUGCAUGCUUUAAGGCGCCCUUUCCACGGGG-------GUGG-AAGCUUGUGAAACUGAAAGUUUUCUUUU ...........(-((((((.((((.(((((....(((((((.....))))-------))))-)))).))))...)))))))......... ( -20.30) >consensus CAGGGAUUCUUU_UUUCACCUGCCUACUUUAAGGCGCCCAUUCAAAAGGAAAAGCAGGAAA_AAACCGGUGAAACCGAAAGUUCCUUGCU ((((((..((((.(((((((.((((......))))..((........))..................)))))))...)))).)))))).. (-16.86 = -17.45 + 0.59)

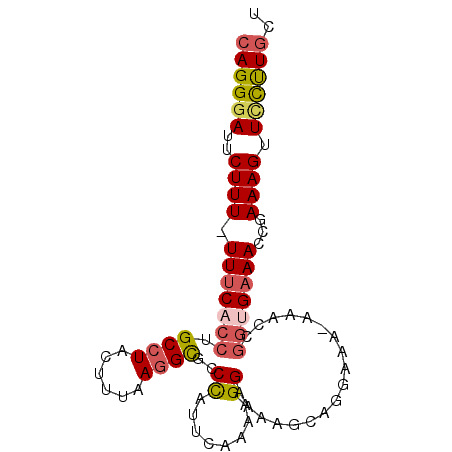

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:31:01 2006