
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 1,445,152 – 1,445,250 |
| Length | 98 |
| Max. P | 0.825577 |

| Location | 1,445,152 – 1,445,250 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 81.12 |
| Mean single sequence MFE | -22.55 |
| Consensus MFE | -17.00 |
| Energy contribution | -16.25 |
| Covariance contribution | -0.75 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.75 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.516358 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
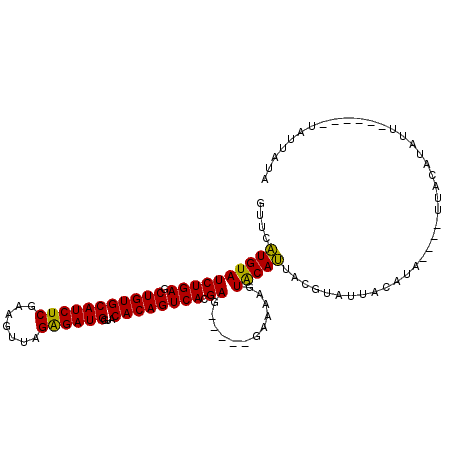
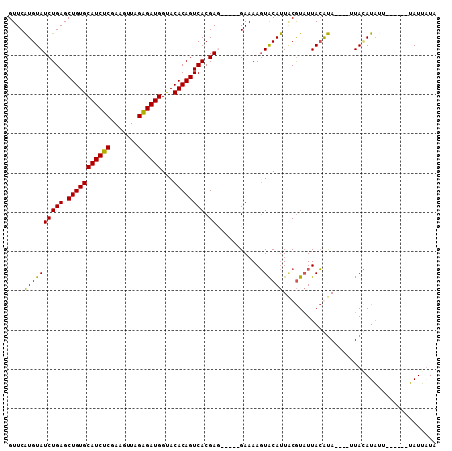
>2L_DroMel_CAF1 1445152 98 + 22407834 GUUCAUGUAUCUGAGCUGUGCAUCUCGAAGUUAGAGAUGGUACACAGUCACGAG-----GAAAAGUACAUUACGUAUUACAUAUUUAUUAUAUUUU------GAUAUUA ....((((((((((.(((((((((((.......))))))...)))))))).)).-----.....((((.....)))))))))...(((((.....)------))))... ( -20.00) >DroSec_CAF1 63175 94 + 1 GUUCAUGUAUCUGAGCUGUGCAUCUCGAAGUUAGAGAUGGUACACAGUCACGAU-----GAAAAGUACAUUACGUAUUACGUA----UUACAUAGU------UAUUAUA .......(((((((.(((((((((((.......))))))...)))))))).)))-----)....(((...((((.....))))----.))).....------....... ( -22.10) >DroSim_CAF1 65693 94 + 1 GUUCAUGUAUCUGAGCUGUGCAUCUCGAAGUUAGAGAUGGUACACAGUCACGAU-----GAAAAGUACAUUGCGUAUUACGUA----UUACAUAUU------UAUUAUA .......(((((((.(((((((((((.......))))))...)))))))).)))-----)....(((...((((.....))))----.))).....------....... ( -22.10) >DroYak_CAF1 64519 105 + 1 GUUCGUGUAUCUGAGCUGUGCAUCUCGAACUUAGGGAUGGCACACAGUCACGAGCCAAGGAAAAUUGCACUCCAUUUUAAACA----UUAAAUAUUUUGCUUUAUUUUG (((((((...(((...((((((((((.......))))).)))))))).))))))).....(((((.(((....((((......----..))))....)))...))))). ( -26.00) >consensus GUUCAUGUAUCUGAGCUGUGCAUCUCGAAGUUAGAGAUGGUACACAGUCACGAG_____GAAAAGUACAUUACGUAUUACAUA____UUACAUAUU______UAUUAUA ....((((((((((.(((((((((((.......))))))...)))))))).))............)))))....................................... (-17.00 = -16.25 + -0.75)

| Location | 1,445,152 – 1,445,250 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 81.12 |
| Mean single sequence MFE | -19.98 |
| Consensus MFE | -14.40 |
| Energy contribution | -13.90 |
| Covariance contribution | -0.50 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.825577 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
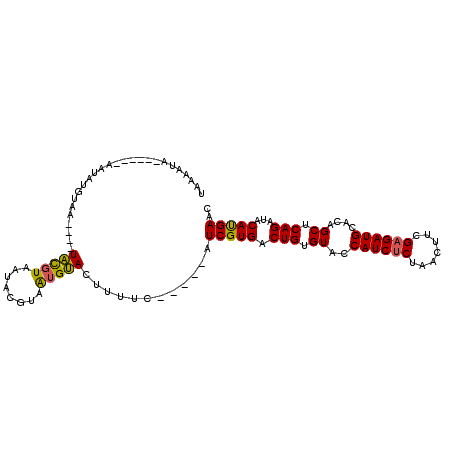
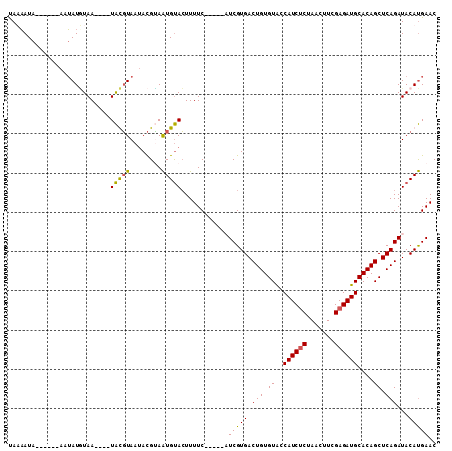
>2L_DroMel_CAF1 1445152 98 - 22407834 UAAUAUC------AAAAUAUAAUAAAUAUGUAAUACGUAAUGUACUUUUC-----CUCGUGACUGUGUACCAUCUCUAACUUCGAGAUGCACAGCUCAGAUACAUGAAC .......------.............((((((.(((.....)))......-----.((.((((((((...((((((.......))))))))))).)))))))))))... ( -18.30) >DroSec_CAF1 63175 94 - 1 UAUAAUA------ACUAUGUAA----UACGUAAUACGUAAUGUACUUUUC-----AUCGUGACUGUGUACCAUCUCUAACUUCGAGAUGCACAGCUCAGAUACAUGAAC .......------..((((((.----((((.....))))...........-----.((.((((((((...((((((.......))))))))))).)))))))))))... ( -21.80) >DroSim_CAF1 65693 94 - 1 UAUAAUA------AAUAUGUAA----UACGUAAUACGCAAUGUACUUUUC-----AUCGUGACUGUGUACCAUCUCUAACUUCGAGAUGCACAGCUCAGAUACAUGAAC .......------..((((((.----(((((........)))))......-----.((.((((((((...((((((.......))))))))))).)))))))))))... ( -20.70) >DroYak_CAF1 64519 105 - 1 CAAAAUAAAGCAAAAUAUUUAA----UGUUUAAAAUGGAGUGCAAUUUUCCUUGGCUCGUGACUGUGUGCCAUCCCUAAGUUCGAGAUGCACAGCUCAGAUACACGAAC .........(((...(((((..----......)))))...))).............(((((.((((((((.(((.(.......).))))))))...)))...))))).. ( -19.10) >consensus UAAAAUA______AAUAUGUAA____UACGUAAUACGUAAUGUACUUUUC_____AUCGUGACUGUGUACCAUCUCUAACUUCGAGAUGCACAGCUCAGAUACAUGAAC ..........................(((((........)))))............(((((.(((.((..((((((.......))))))....)).)))...))))).. (-14.40 = -13.90 + -0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:37:20 2006