
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,578,606 – 13,578,722 |
| Length | 116 |
| Max. P | 0.515123 |

| Location | 13,578,606 – 13,578,722 |
|---|---|
| Length | 116 |
| Sequences | 4 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 83.38 |
| Mean single sequence MFE | -31.28 |
| Consensus MFE | -18.76 |
| Energy contribution | -21.32 |
| Covariance contribution | 2.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.60 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.515123 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
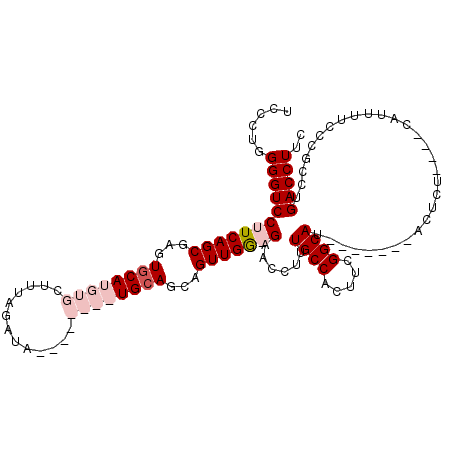
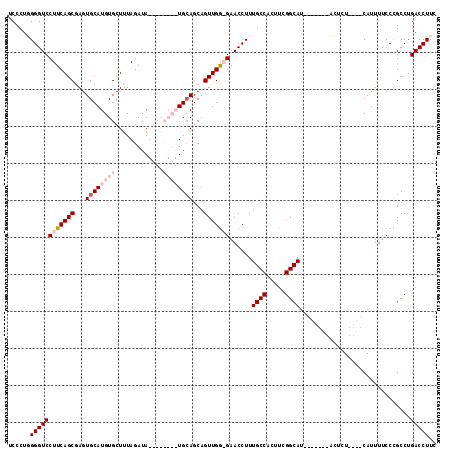
>2L_DroMel_CAF1 13578606 116 + 22407834 UCCCUUGGGUCCUUCAGCGAGUGCAUGUGCUUUAGAUAUGUACAUAUGGAGCAGUUGGGGAACCUUUGCCACUUCGGCAUGGAGGAGAGUAUCCUCCAUUUUCCCGCCUGACCUUC ......((((((((((....((((((((.......))))))))...))))).((.(((((((....((((.....))))(((((((.....))))))).))))))).))))))).. ( -42.80) >DroSec_CAF1 11755 109 + 1 UCCCUUGGGUCCUUCAGCGAGUGCAUGUGCUUUAGAUAUGUACAUAUGCAGCAGUUGGGGAACCUUUGCCACUUCGGCAU-------ACUCUCCUCCAUUUUCCCGCCUGACCUUC ......(((((.....(((..((((((((.............)))))))).....((((((.....((((.....)))).-------....))).)))......)))..))))).. ( -29.12) >DroEre_CAF1 12128 96 + 1 UCCCUGGGGUCCUUCAGCAAGUGCAUGUGCUUUAGAUA--------UGCAGCAGUUGA-GAACCUUUGCCUCUUCGGCAU-------CCUCU----CAUUUUCCUGCCUGACCUUC .....((((((..........(((((((.......)))--------))))((((.(((-((.....((((.....)))).-------..)))----)).....))))..)))))). ( -27.10) >DroYak_CAF1 8893 96 + 1 UCCCUGGGGUCCUUCAGCGAGUGCAUGUGCUUUAGAUA--------UGCAGCUGUUGG-GAACCUUUGCCUCUUCGGCAU-------CCUCU----CAUUUUCCUGCCUGACCUUU .....((((((...(((.((((((((((.......)))--------)))).....(((-((.....((((.....)))).-------..)))----))..))))))...)))))). ( -26.10) >consensus UCCCUGGGGUCCUUCAGCGAGUGCAUGUGCUUUAGAUA________UGCAGCAGUUGG_GAACCUUUGCCACUUCGGCAU_______ACUCU____CAUUUUCCCGCCUGACCUUC ......((((((((((((...((((((((.............))))))))...)))))))......((((.....))))..............................))))).. (-18.76 = -21.32 + 2.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:26:42 2006