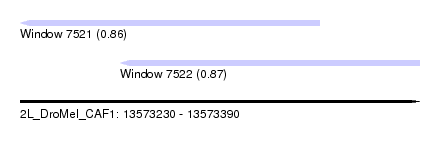
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,573,230 – 13,573,390 |
| Length | 160 |
| Max. P | 0.870948 |

| Location | 13,573,230 – 13,573,350 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 93.47 |
| Mean single sequence MFE | -35.73 |
| Consensus MFE | -32.14 |
| Energy contribution | -31.82 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.856134 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13573230 120 - 22407834 ACAAUGGGCGCUAGGGGGUGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAAGUAAAUGACGACUGCUAAAUGUGUCAAUUUUGCGACACGA .........((((((((((.((((.(((.((.....)).))).)))).)))).)).))))(((((.((..(((.((....))..)))..)))))))...((((((........)))))). ( -37.70) >DroSec_CAF1 6471 120 - 1 ACAAUGGGCGCUAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAAGUAAAUGACGACUGCUAAAUGUGUCAAUUCUGCGACACGA .........(((((((((..((((.(((.((.....)).))).))))..))).)).))))(((((.((..(((.((....))..)))..)))))))...((((((........)))))). ( -36.70) >DroSim_CAF1 6048 120 - 1 ACAAUGGGCGCCAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAAGUAAAUGACGACUGCUAAAUGUGUCAAUUCUGCGACACGA .........(((((((((..((((.(((.((.....)).))).))))..))).)).))))(((((.((..(((.((....))..)))..)))))))...((((((........)))))). ( -39.00) >DroYak_CAF1 3580 119 - 1 GCAAUGGGCGACAAGGGGCGUGGCAGUUGCCAGAA-GGGAGUGGCUGGAUCCACCAUGGCAGCAGAUUAGCAUAACAAAAGUAAAUGACGACUGCUAAAUGUGUCAGUUUUACGAAGCGA ((.(((.((.((.......)).)).(((((((...-..(.((((......))))).))))))).......))).......(((((((((.((........))))))..)))))...)).. ( -29.50) >consensus ACAAUGGGCGCCAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAAGUAAAUGACGACUGCUAAAUGUGUCAAUUCUGCGACACGA .........((((.((((..((((..((.((.....)).))..))))...)).)).))))(((((.((..(((.((....))..)))..)))))))...((((((........)))))). (-32.14 = -31.82 + -0.31)


| Location | 13,573,270 – 13,573,390 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.86 |
| Mean single sequence MFE | -38.34 |
| Consensus MFE | -34.33 |
| Energy contribution | -34.95 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.90 |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.870948 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13573270 120 - 22407834 CAGCAAAGCGUUAUAAUAGGGCUUAUUAGGGUUACUCCCGACAAUGGGCGCUAGGGGGUGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAA .........(((((......(((((((.(((.....)))...)))))))((((((((((.((((.(((.((.....)).))).)))).)))).)).)))).((......))))))).... ( -39.60) >DroSec_CAF1 6511 120 - 1 CAGCAAAGCGUUAUAAUAGGGCUUAUUAGGGUUACCCCCGACAAUGGGCGCUAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAA ..((...(((((....(((.(((((((.(((.....)))...))))))).)))...))))).)).(((((((.....(((.........)))....)))))))................. ( -40.10) >DroSim_CAF1 6088 120 - 1 CAGCAAAGCGUUAUAAUAGGGCUUAUUAGGGUUACCCCCGACAAUGGGCGCCAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAA .........(((((......(((((((.(((.....)))...)))))))(((((((((..((((.(((.((.....)).))).))))..))).)).)))).((......))))))).... ( -40.40) >DroYak_CAF1 3620 118 - 1 CAGCAAAGCGUUAUAAUAGGGCUUAUUAGGGUUACC-CCGGCAAUGGGCGACAAGGGGCGUGGCAGUUGCCAGAA-GGGAGUGGCUGGAUCCACCAUGGCAGCAGAUUAGCAUAACAAAA ..((...(((((........(((((((.(((....)-))...))))))).......))))).)).(((((((...-..(.((((......))))).)))))))................. ( -33.26) >consensus CAGCAAAGCGUUAUAAUAGGGCUUAUUAGGGUUACCCCCGACAAUGGGCGCCAGGGGGCGUGGCAGCUGCCAAAAAGGGAGUGGCUAAAUCCACCAUGGCAGCAGAUUAGCAUAACAAAA ..((...(((((....(((.(((((((.(((.....)))...))))))).)))...))))).)).(((((((.....(((.........)))....)))))))................. (-34.33 = -34.95 + 0.62)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:26:25 2006