
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,554,956 – 13,555,051 |
| Length | 95 |
| Max. P | 0.998663 |

| Location | 13,554,956 – 13,555,051 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 94.53 |
| Mean single sequence MFE | -22.92 |
| Consensus MFE | -20.64 |
| Energy contribution | -21.24 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.19 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.18 |
| SVM RNA-class probability | 0.998663 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
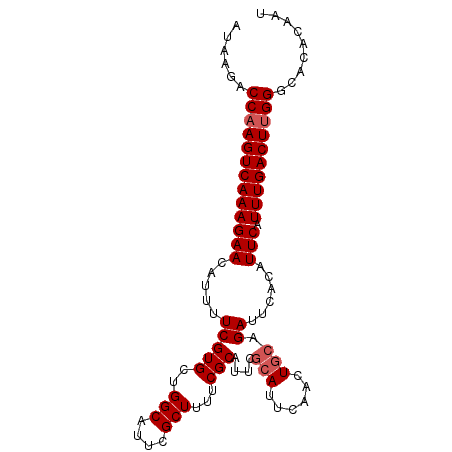
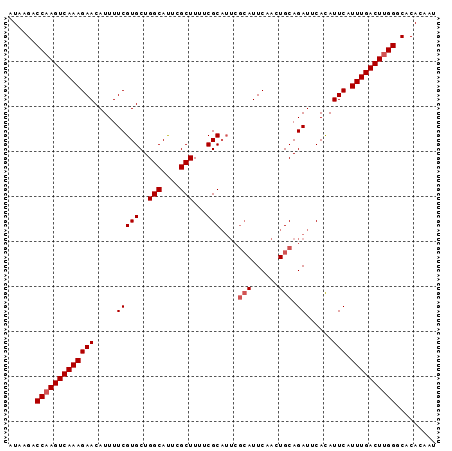
>2L_DroMel_CAF1 13554956 95 + 22407834 AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCAUUCGCUUUUCGCAUUCGCAUUCAACUGCAGAUUCACAUUCAUUUGACUCGGGCACACAAU ......((.((((((((((.....(((((..(((....)))...)))....(((......))).))......))).))))))).))......... ( -19.50) >DroSec_CAF1 16239 95 + 1 AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCAUUUGCUUUUCGCAUUCGCAUUCAACUGCAGAUUCACAUUCAUUUGACUUGGGCACACAAU ......(((((((((((((.....(((((..(((....)))...)))....(((......))).))......))).))))))))))......... ( -23.90) >DroSim_CAF1 23745 95 + 1 AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCAUUCGCUUUUCGCAUUCGCAUUCAACUGCAGAUUCAUAUUCAUUUGACUUGGGCACACAAU ......(((((((((((((.((..(((((..(((....)))...)))....(((......))).))...)).))).))))))))))......... ( -23.80) >DroEre_CAF1 16184 95 + 1 AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCACUCGCUUUUCGCAUUCGCAUUCAAGUGCAGAUUCACAUUCAUUUGACUUGGGCACAGAAU ......(((((((((((((.....(((((..(((....)))...)))....((((....)))).))......))).))))))))))......... ( -25.10) >DroYak_CAF1 14419 89 + 1 AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCAUUCGCUUUUCGC------AUUCAAGUGCAGAUUCACAUUCAUUUGACUUGGGCACAGAAU ......(((((((((((((.......(((..(((....)))...)))------......(((......))).))).))))))))))......... ( -22.30) >consensus AUAAGACCAAGUCAAAGAACAUUUUCGUGCUGGCAUUCGCUUUUCGCAUUCGCAUUCAACUGCAGAUUCACAUUCAUUUGACUUGGGCACACAAU ......(((((((((((((.....(((((..(((....)))...)))....(((......))).))......))).))))))))))......... (-20.64 = -21.24 + 0.60)

| Location | 13,554,956 – 13,555,051 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 94.53 |
| Mean single sequence MFE | -25.33 |
| Consensus MFE | -22.82 |
| Energy contribution | -23.18 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.32 |
| SVM RNA-class probability | 0.992296 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
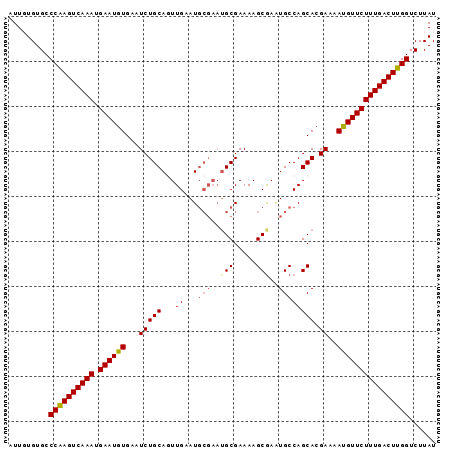
>2L_DroMel_CAF1 13554956 95 - 22407834 AUUGUGUGCCCGAGUCAAAUGAAUGUGAAUCUGCAGUUGAAUGCGAAUGCGAAAAGCGAAUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((...((((((.(((....))).))))....((........))..))..))))))))))))))))...... ( -24.20) >DroSec_CAF1 16239 95 - 1 AUUGUGUGCCCAAGUCAAAUGAAUGUGAAUCUGCAGUUGAAUGCGAAUGCGAAAAGCAAAUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((...(((((.((.....))...(((.....))).......))).))..))))))))))))))))...... ( -25.10) >DroSim_CAF1 23745 95 - 1 AUUGUGUGCCCAAGUCAAAUGAAUAUGAAUCUGCAGUUGAAUGCGAAUGCGAAAAGCGAAUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((...((((((.(((....))).))))....((........))..))..))))))))))))))))...... ( -25.50) >DroEre_CAF1 16184 95 - 1 AUUCUGUGCCCAAGUCAAAUGAAUGUGAAUCUGCACUUGAAUGCGAAUGCGAAAAGCGAGUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((....(((((((((....((....)).....)))))).))).......))))))))))))))))...... ( -28.50) >DroYak_CAF1 14419 89 - 1 AUUCUGUGCCCAAGUCAAAUGAAUGUGAAUCUGCACUUGAAU------GCGAAAAGCGAAUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((...(((((.......(------((.....))).......))).))..))))))))))))))))...... ( -23.34) >consensus AUUGUGUGCCCAAGUCAAAUGAAUGUGAAUCUGCAGUUGAAUGCGAAUGCGAAAAGCGAAUGCCAGCACGAAAAUGUUCUUUGACUUGGUCUUAU .........((((((((((.((((((...(((((...((...(((..(((.....)))..)))))))).))..))))))))))))))))...... (-22.82 = -23.18 + 0.36)
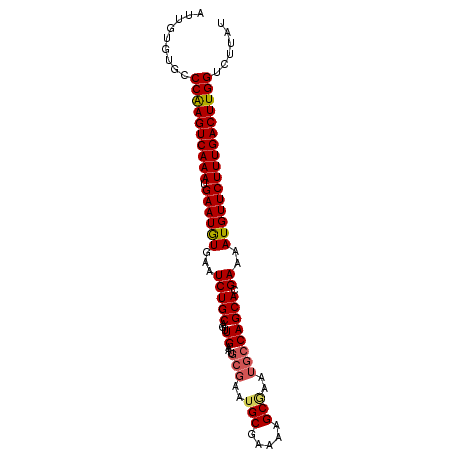


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:25:50 2006