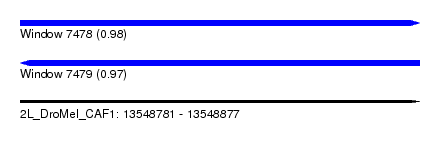
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,548,781 – 13,548,877 |
| Length | 96 |
| Max. P | 0.982894 |

| Location | 13,548,781 – 13,548,877 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | forward |
| Mean pairwise identity | 89.95 |
| Mean single sequence MFE | -33.30 |
| Consensus MFE | -27.88 |
| Energy contribution | -28.00 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.93 |
| SVM RNA-class probability | 0.982894 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13548781 96 + 22407834 AUCGAAUGAAAAGAAGAAGAAAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCAGCAGUUGAGCUUCUAGU------- ..............(((((....(((.(.((.(.((((((((((..((.((........---))..)).)))))))))).))).).)))..)))))...------- ( -30.30) >DroSec_CAF1 7641 103 + 1 AUCGAAAGAAGAGAAGAAGAAAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCAGCAGUUGAGCUUCUAGUUUCGCAU .((....)).(((((((((....(((.(.((.(.((((((((((..((.((........---))..)).)))))))))).))).).)))..)))))..)))).... ( -34.80) >DroSim_CAF1 7868 103 + 1 AUCGAAAGAAGAGAAGAAGAAAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU---GUAGUGACCACUGUGCGGCAGCAGUUGAGCUUCUAGUUUCGCAU .((....)).(((((((((....(((.(.((.(.((((((((((..((.((........---))..)).)))))))))).))).).)))..)))))..)))).... ( -34.80) >DroEre_CAF1 9982 106 + 1 GUCAAAAGAAGAGAUGGAGAAAACAAGUUCUGGGCGCACAGUGGUGCAAGCUUAGAAGUCUAGUGGUGACCACUGUGCGGCAGCAGUUAAGCUUCUAGAUUCGCAU .......(((....((((..............(.(((((((((((.((.(((.((....)))))..))))))))))))).)(((......)))))))..))).... ( -33.30) >consensus AUCGAAAGAAGAGAAGAAGAAAACAAGUUCUGGGCGCACAGUGGGGCAAGCUUAGAACU___GUAGUGACCACUGUGCGGCAGCAGUUGAGCUUCUAGUUUCGCAU .((....)).....(((((....(((.(.((.(.((((((((((.....(((............)))..)))))))))).))).).)))..))))).......... (-27.88 = -28.00 + 0.13)


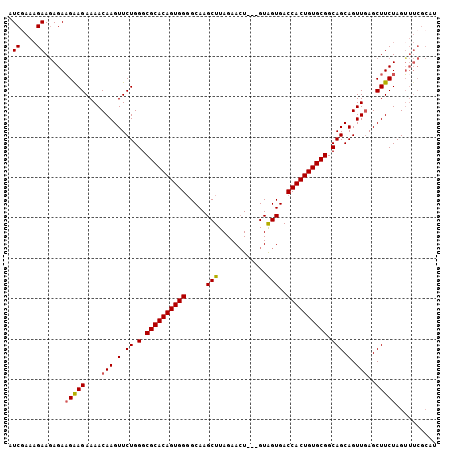
| Location | 13,548,781 – 13,548,877 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 89.95 |
| Mean single sequence MFE | -26.80 |
| Consensus MFE | -22.17 |
| Energy contribution | -21.98 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.54 |
| Structure conservation index | 0.83 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968891 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
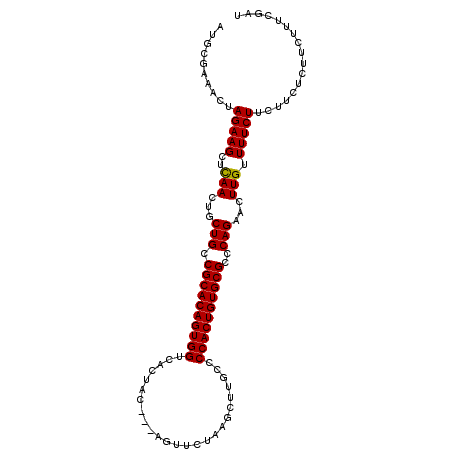
>2L_DroMel_CAF1 13548781 96 - 22407834 -------ACUAGAAGCUCAACUGCUGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUUUCUUCUUCUUUUCAUUCGAU -------...(((((..(((...(((.((((((((((.((....---(((.....)))))..))))))))))..)))...)))....))))).............. ( -24.90) >DroSec_CAF1 7641 103 - 1 AUGCGAAACUAGAAGCUCAACUGCUGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUUUCUUCUUCUCUUCUUUCGAU ...(((((..(((((..(((...(((.((((((((((.((....---(((.....)))))..))))))))))..)))...)))....))))).......))))).. ( -27.40) >DroSim_CAF1 7868 103 - 1 AUGCGAAACUAGAAGCUCAACUGCUGCCGCACAGUGGUCACUAC---AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUUUCUUCUUCUCUUCUUUCGAU ...(((((..(((((..(((...(((.((((((((((.((....---(((.....)))))..))))))))))..)))...)))....))))).......))))).. ( -27.40) >DroEre_CAF1 9982 106 - 1 AUGCGAAUCUAGAAGCUUAACUGCUGCCGCACAGUGGUCACCACUAGACUUCUAAGCUUGCACCACUGUGCGCCCAGAACUUGUUUUCUCCAUCUCUUCUUUUGAC ...((((...((((((......)))..(((((((((((((....(((....)))....)).)))))))))))......................)))...)))).. ( -27.50) >consensus AUGCGAAACUAGAAGCUCAACUGCUGCCGCACAGUGGUCACUAC___AGUUCUAAGCUUGCCCCACUGUGCGCCCAGAACUUGUUUUCUUCUUCUCUUCUUUCGAU ..........(((((..(((...(((.((((((((((.........................))))))))))..)))...))).)))))................. (-22.17 = -21.98 + -0.19)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:25:42 2006