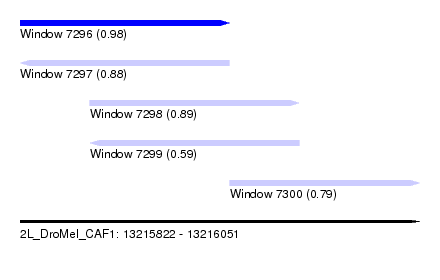
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,215,822 – 13,216,051 |
| Length | 229 |
| Max. P | 0.983853 |

| Location | 13,215,822 – 13,215,942 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 97.08 |
| Mean single sequence MFE | -30.25 |
| Consensus MFE | -30.00 |
| Energy contribution | -30.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.99 |
| SVM decision value | 1.96 |
| SVM RNA-class probability | 0.983853 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
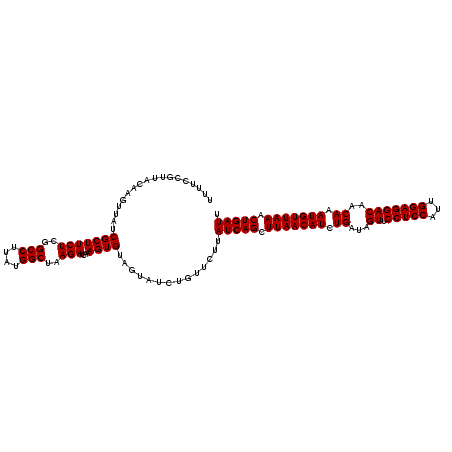
>2L_DroMel_CAF1 13215822 120 + 22407834 UUUUCCGUUUCAAGUUAUCGCUUCUCGGCCUUAUGGCUAAGAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUU ............((.((((((((((..(((....)))..)))....))))..))).))......(((((.(((((((.((...((.(((((...)))))))..)).))))))).))))). ( -30.30) >DroSec_CAF1 3047 120 + 1 UUUUUCGUUUCAAGUUAUCGCUUCUCGGCCUUAUGGCUAAGAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUU ............((.((((((((((..(((....)))..)))....))))..))).))......(((((.(((((((.((...((.(((((...)))))))..)).))))))).))))). ( -30.30) >DroEre_CAF1 3276 120 + 1 UUUUCCCUUACAAUUUAUCGCUUCUCGGCCUUAUGGCUAAGAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUU .........(((......(((((((..(((....)))..)))....)))).......)))....(((((.(((((((.((...((.(((((...)))))))..)).))))))).))))). ( -30.12) >DroYak_CAF1 7908 119 + 1 UUUUCGUUUA-AAGUUAUCGCUUCUCGGCCUUAUGGCUAAGAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUU ..........-.((.((((((((((..(((....)))..)))....))))..))).))......(((((.(((((((.((...((.(((((...)))))))..)).))))))).))))). ( -30.30) >consensus UUUUCCGUUACAAGUUAUCGCUUCUCGGCCUUAUGGCUAAGAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUU ..................(((((((..(((....)))..)))....))))..............(((((.(((((((.((...((.(((((...)))))))..)).))))))).))))). (-30.00 = -30.00 + 0.00)

| Location | 13,215,822 – 13,215,942 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 97.08 |
| Mean single sequence MFE | -38.88 |
| Consensus MFE | -38.30 |
| Energy contribution | -38.30 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -5.30 |
| Structure conservation index | 0.99 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.879399 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
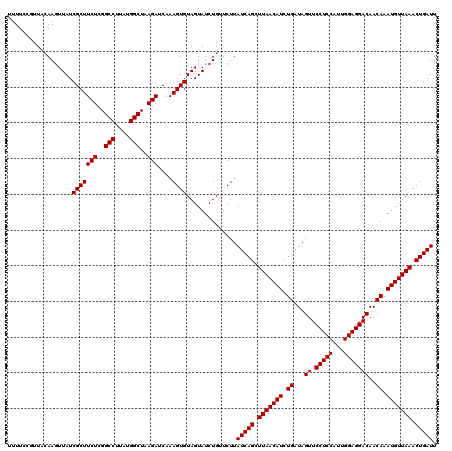
>2L_DroMel_CAF1 13215822 120 - 22407834 AAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUCUUAGCCAUAAGGCCGAGAAGCGAUAACUUGAAACGGAAAA .((((((((((((((((...((((((...)))))).....))))))))))))))))(((..((((........)))).(((..(((....)))..)))........)))........... ( -38.70) >DroSec_CAF1 3047 120 - 1 AAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUCUUAGCCAUAAGGCCGAGAAGCGAUAACUUGAAACGAAAAA .((((((((((((((((...((((((...)))))).....))))))))))))))))(((..((((........)))).(((..(((....)))..)))........)))........... ( -38.70) >DroEre_CAF1 3276 120 - 1 AAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUCUUAGCCAUAAGGCCGAGAAGCGAUAAAUUGUAAGGGAAAA .((((((((((((((((...((((((...)))))).....)))))))))))))))).........(((......(((.(((..(((....)))..)))..)))......)))........ ( -39.80) >DroYak_CAF1 7908 119 - 1 AAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUCUUAGCCAUAAGGCCGAGAAGCGAUAACUU-UAAACGAAAA .((((((((((((((((...((((((...)))))).....)))))))))))))))).....((((........)))).(((..(((....)))..)))...........-.......... ( -38.30) >consensus AAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUCUUAGCCAUAAGGCCGAGAAGCGAUAACUUGAAACGGAAAA .((((((((((((((((...((((((...)))))).....)))))))))))))))).....((((........)))).(((..(((....)))..)))...................... (-38.30 = -38.30 + 0.00)

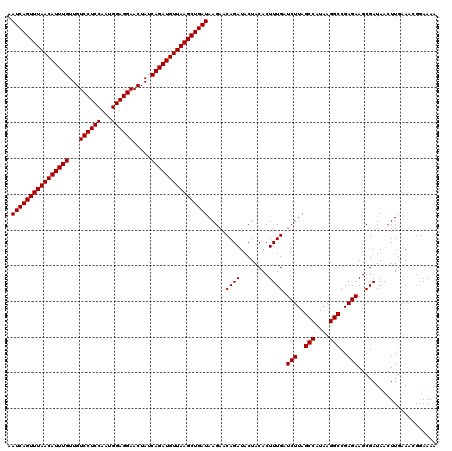
| Location | 13,215,862 – 13,215,982 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -35.40 |
| Consensus MFE | -35.40 |
| Energy contribution | -35.40 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.71 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.893023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
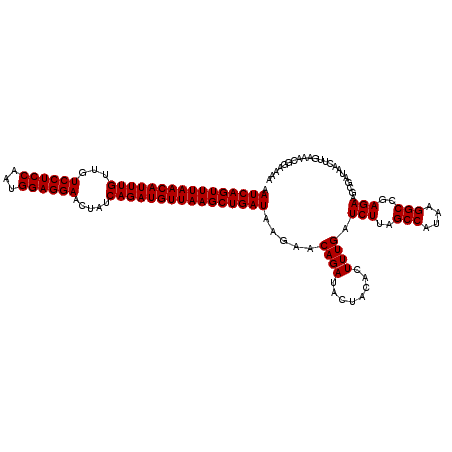
>2L_DroMel_CAF1 13215862 120 + 22407834 GAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUUUUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCU ......((..(....((((((((.(((((.(((((((.((...((.(((((...)))))))..)).))))))).)))))....)))).))))....)..)).((((....))))...... ( -35.40) >DroSec_CAF1 3087 120 + 1 GAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUUUUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCU ......((..(....((((((((.(((((.(((((((.((...((.(((((...)))))))..)).))))))).)))))....)))).))))....)..)).((((....))))...... ( -35.40) >DroEre_CAF1 3316 120 + 1 GAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUUUUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCU ......((..(....((((((((.(((((.(((((((.((...((.(((((...)))))))..)).))))))).)))))....)))).))))....)..)).((((....))))...... ( -35.40) >DroYak_CAF1 7947 120 + 1 GAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUUUUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCU ......((..(....((((((((.(((((.(((((((.((...((.(((((...)))))))..)).))))))).)))))....)))).))))....)..)).((((....))))...... ( -35.40) >consensus GAUCAAAGUGUAGUAUCUGUUCUUAUCAGCUUAACAUCUGAUAGUUCCUCCAUUGGAGGACAACAAAUGUUAAACUGAUUUUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCU ......((..(....((((((((.(((((.(((((((.((...((.(((((...)))))))..)).))))))).)))))....)))).))))....)..)).((((....))))...... (-35.40 = -35.40 + -0.00)
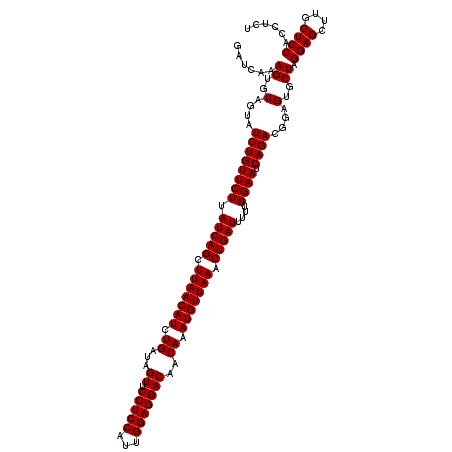


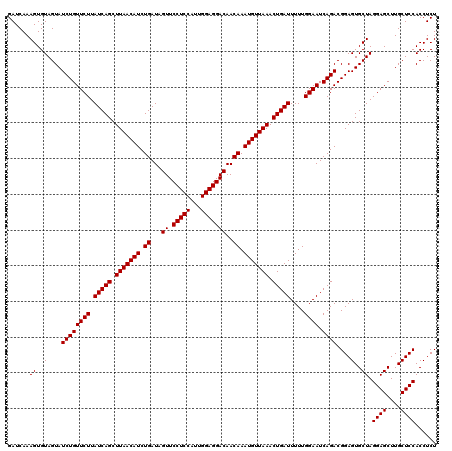
| Location | 13,215,862 – 13,215,982 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 100.00 |
| Mean single sequence MFE | -43.60 |
| Consensus MFE | -43.60 |
| Energy contribution | -43.60 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -5.57 |
| Structure conservation index | 1.00 |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.594800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13215862 120 - 22407834 AGAGGUGGAGCAAGCUCCUAGCACUCCGUCUGAUUCCAAAAAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUC .(((((((((...((.....)).))))(((((.(((.....((((((((((((((((...((((((...)))))).....))))))))))))))))..)))))))).....))))).... ( -43.60) >DroSec_CAF1 3087 120 - 1 AGAGGUGGAGCAAGCUCCUAGCACUCCGUCUGAUUCCAAAAAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUC .(((((((((...((.....)).))))(((((.(((.....((((((((((((((((...((((((...)))))).....))))))))))))))))..)))))))).....))))).... ( -43.60) >DroEre_CAF1 3316 120 - 1 AGAGGUGGAGCAAGCUCCUAGCACUCCGUCUGAUUCCAAAAAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUC .(((((((((...((.....)).))))(((((.(((.....((((((((((((((((...((((((...)))))).....))))))))))))))))..)))))))).....))))).... ( -43.60) >DroYak_CAF1 7947 120 - 1 AGAGGUGGAGCAAGCUCCUAGCACUCCGUCUGAUUCCAAAAAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUC .(((((((((...((.....)).))))(((((.(((.....((((((((((((((((...((((((...)))))).....))))))))))))))))..)))))))).....))))).... ( -43.60) >consensus AGAGGUGGAGCAAGCUCCUAGCACUCCGUCUGAUUCCAAAAAUCAGUUUAACAUUUGUUGUCCUCCAAUGGAGGAACUAUCAGAUGUUAAGCUGAUAAGAACAGAUACUACACUUUGAUC .(((((((((...((.....)).))))(((((.(((.....((((((((((((((((...((((((...)))))).....))))))))))))))))..)))))))).....))))).... (-43.60 = -43.60 + 0.00)

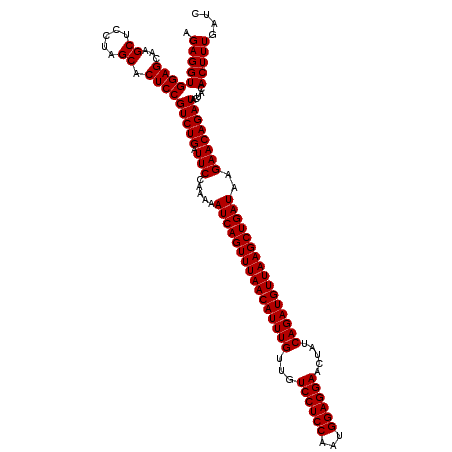
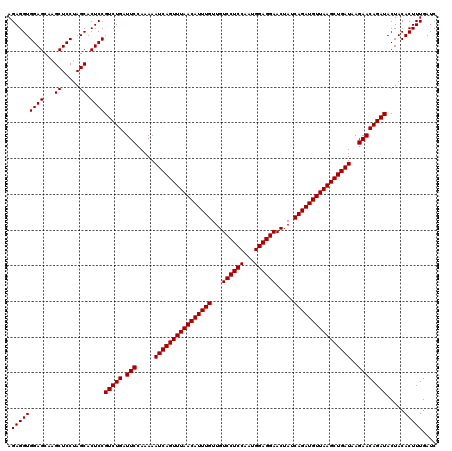
| Location | 13,215,942 – 13,216,051 |
|---|---|
| Length | 109 |
| Sequences | 3 |
| Columns | 109 |
| Reading direction | forward |
| Mean pairwise identity | 97.55 |
| Mean single sequence MFE | -36.97 |
| Consensus MFE | -35.89 |
| Energy contribution | -35.67 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.81 |
| Structure conservation index | 0.97 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.792535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13215942 109 + 22407834 UUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCUGUCGCGGGUUGGCCCGGUAUUGCAGUACCGCCGGGAUUUCGGCCCAACUUAAUAAUAAAUAUAAAACCA ..(((.....(((((((.(...((((....)))).))))))))..(((((((((((((...........))))))...))))))).....................))) ( -37.30) >DroSec_CAF1 3167 109 + 1 UUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCUGUCGCGGGUUGGCCCGGUAUUGCAGUACCGCCGGGAUUUCGGCCCAACUGAAUAAUAAAUAUAAAGCUA ((..(.....(((((((.(...((((....)))).))))))))..(((((((((((((...........))))))...)))))))..)..))................. ( -36.70) >DroEre_CAF1 3396 109 + 1 UUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCUGUCGCGGGUUGGCCCGGUAUUGCAGUACCGCCGGGAUUUCGGCCCAACUGAAAAAUAAAUAUAAAACUA ((..(.....(((((((.(...((((....)))).))))))))..(((((((((((((...........))))))...)))))))..)..))................. ( -36.90) >consensus UUUGGAAUCAGACGGAGUGCUAGGAGCUUGCUCCACCUCUGUCGCGGGUUGGCCCGGUAUUGCAGUACCGCCGGGAUUUCGGCCCAACUGAAUAAUAAAUAUAAAACUA ..(((.....(((((((.(...((((....)))).))))))))..(((((((((((((...........))))))...))))))).....................))) (-35.89 = -35.67 + -0.22)

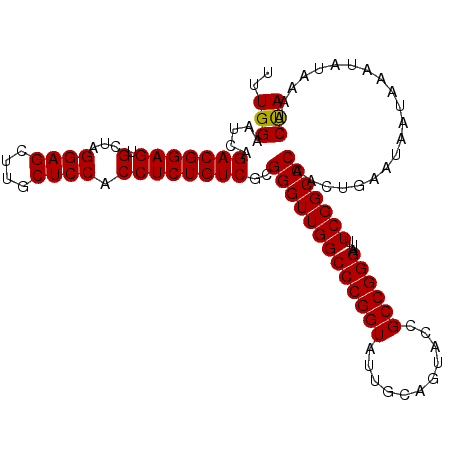
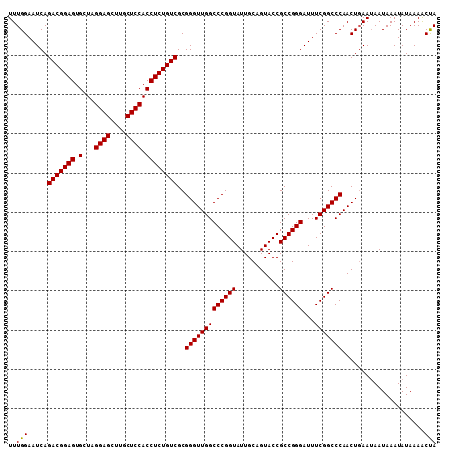
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:51 2006