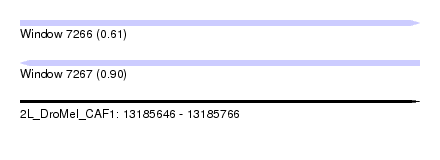

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,185,646 – 13,185,766 |
| Length | 120 |
| Max. P | 0.898281 |

| Location | 13,185,646 – 13,185,766 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 79.45 |
| Mean single sequence MFE | -44.10 |
| Consensus MFE | -27.93 |
| Energy contribution | -29.97 |
| Covariance contribution | 2.03 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.606175 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13185646 120 + 22407834 CCUGCAUCACAGGAGCAACGGUUGAGGUCAGCGAAGUGGUCUCCACGGGAUUUAUGCGAAGCAUUUGGUAUGGUGCUUCGCCCUCCUUCUGCAACUCAUGCAGUGUGCUACCAUCUCGCU ((((.....))))(((....((((....))))((..((((...(((((((.....((((((((((......))))))))))..)))).(((((.....))))).)))..))))..))))) ( -42.90) >DroVir_CAF1 6439 105 + 1 UCU-----------GCAACG----UCGUCAAUGUCGACUCGUCGACUGGAUUGAGAUGCACCAUCUGAUGUGGUGAUCCGUCUUCGGUGAGCAGCUCGUGCAGCGUGCUACCAUCUUGCU ...-----------((((((----((.((((((((((....)))))...)))))))))((((((.....))))))..........(((.(((((((.....))).)))))))...)))). ( -32.90) >DroSec_CAF1 6106 115 + 1 -----AUCGCAGGAGCAACGGAAGAGGUCAGCGAAGUACUCUCCACCGGGUUGAGGCGAAGCAUUUGGUAGGGUGCUUCGCCCUCCUUCUGCAGCUCAUGCAGUGUGCUACCAUCUCGCU -----...(((.((((..(((((((((...(((((((((((((((....(((.......)))...))).))))))))))))))).))))))..)))).)))((((...........)))) ( -46.50) >DroSim_CAF1 6081 115 + 1 -----AUCGCAGGAGCAACGGAUGAGGUCAGCGAAGUGGUCUCCACCGGAUUUAGGCGAAGCAUUUGGUAGGGUGCUUCGCCCUCCUUCUGCAGCUCAUGCAGUGUGCUACCAUCUCGCU -----...(((((((...(((..(((..((......))..)))..)))......((((((((((((....))))))))))))...)))))))(((.(((...))).)))........... ( -43.80) >DroEre_CAF1 6202 120 + 1 GCUGUAUCGCAGCAGUAACGGAGGAGGCCAGCGAAGUGGUCUCCACGGGAUUUAGGCGAAGCAUUUGGUAGGGUGCUUCGCCCUCCUUCUGCAGCUCAUGAAGAGUGCUACCAUCUCGCU ((((.....)))).........((((((((......)))))))).((((((...((((((((((((....))))))))))))........(((.(((.....))))))....)))))).. ( -49.60) >DroYak_CAF1 6182 120 + 1 CCUGCAUCGCAGGUGCAACGGAGGAGGUCAGCGAAGUGGUCUCCACGGGAUUUAGGCGAAGCAUUUGGUAGGGUGCUUCGCCCUCCUUCUGCAGCUCAUGGAGAGUGCUACCAUCGCGCU .((.(((.(.((.((((..(((((((.........((((...))))........((((((((((((....))))))))))))))))))))))).)))))).))(((((.......))))) ( -48.90) >consensus _CU__AUCGCAGGAGCAACGGAGGAGGUCAGCGAAGUGGUCUCCACGGGAUUUAGGCGAAGCAUUUGGUAGGGUGCUUCGCCCUCCUUCUGCAGCUCAUGCAGUGUGCUACCAUCUCGCU .............................(((((.(((((...(((.(((....(((((((((((......))))))))))).)))..(((((.....))))).)))..))))).))))) (-27.93 = -29.97 + 2.03)



| Location | 13,185,646 – 13,185,766 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 79.45 |
| Mean single sequence MFE | -43.32 |
| Consensus MFE | -26.80 |
| Energy contribution | -29.80 |
| Covariance contribution | 3.00 |
| Combinations/Pair | 1.19 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.898281 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 13185646 120 - 22407834 AGCGAGAUGGUAGCACACUGCAUGAGUUGCAGAAGGAGGGCGAAGCACCAUACCAAAUGCUUCGCAUAAAUCCCGUGGAGACCACUUCGCUGACCUCAACCGUUGCUCCUGUGAUGCAGG ((((((.((((..(((.(((((.....)))))..(((..((((((((..........)))))))).....))).)))...)))).))))))..(((....(((..(....)..))).))) ( -43.40) >DroVir_CAF1 6439 105 - 1 AGCAAGAUGGUAGCACGCUGCACGAGCUGCUCACCGAAGACGGAUCACCACAUCAGAUGGUGCAUCUCAAUCCAGUCGACGAGUCGACAUUGACGA----CGUUGC-----------AGA .((....((((((((.(((.....))))))).))))..(((((((((((((....).))))).)))(((((...(((((....))))))))))...----))))))-----------... ( -33.00) >DroSec_CAF1 6106 115 - 1 AGCGAGAUGGUAGCACACUGCAUGAGCUGCAGAAGGAGGGCGAAGCACCCUACCAAAUGCUUCGCCUCAACCCGGUGGAGAGUACUUCGCUGACCUCUUCCGUUGCUCCUGCGAU----- .((.........))....((((.((((.((.((((..((((((((((..........)))))))))......((((((((....)))))))).)..)))).)).)))).))))..----- ( -44.30) >DroSim_CAF1 6081 115 - 1 AGCGAGAUGGUAGCACACUGCAUGAGCUGCAGAAGGAGGGCGAAGCACCCUACCAAAUGCUUCGCCUAAAUCCGGUGGAGACCACUUCGCUGACCUCAUCCGUUGCUCCUGCGAU----- ((((((.((((..(((.(((((.....)))))..(((((((((((((..........))))))))))...))).)))...)))).))))))..........(((((....)))))----- ( -45.30) >DroEre_CAF1 6202 120 - 1 AGCGAGAUGGUAGCACUCUUCAUGAGCUGCAGAAGGAGGGCGAAGCACCCUACCAAAUGCUUCGCCUAAAUCCCGUGGAGACCACUUCGCUGGCCUCCUCCGUUACUGCUGCGAUACAGC .((...((.(((((((((.....))).....(.((((((((((((((..........))))))))).....((.((((((....)))))).)).))))).).....)))))).))...)) ( -43.80) >DroYak_CAF1 6182 120 - 1 AGCGCGAUGGUAGCACUCUCCAUGAGCUGCAGAAGGAGGGCGAAGCACCCUACCAAAUGCUUCGCCUAAAUCCCGUGGAGACCACUUCGCUGACCUCCUCCGUUGCACCUGCGAUGCAGG ...(((((((..((((((.....))).)))...((((((((((((((..........)))))))))........((((((....))))))....)))))))))))).(((((...))))) ( -50.10) >consensus AGCGAGAUGGUAGCACACUGCAUGAGCUGCAGAAGGAGGGCGAAGCACCCUACCAAAUGCUUCGCCUAAAUCCCGUGGAGACCACUUCGCUGACCUCAUCCGUUGCUCCUGCGAU__AG_ ((((((.((((..(((.(((((.....)))))..(((((((((((((..........))))))))))...))).)))...)))).))))))..........(((((....)))))..... (-26.80 = -29.80 + 3.00)
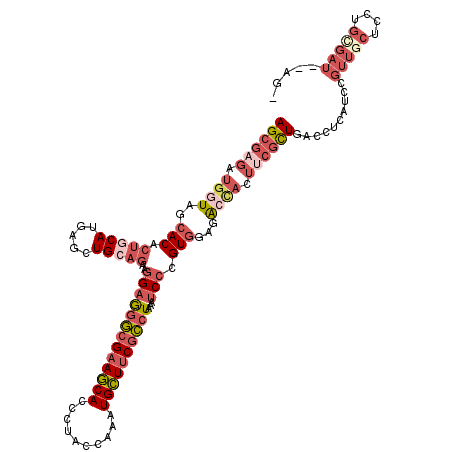


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:22:20 2006