
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 13,133,540 – 13,133,655 |
| Length | 115 |
| Max. P | 0.740565 |

| Location | 13,133,540 – 13,133,655 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 82.17 |
| Mean single sequence MFE | -48.18 |
| Consensus MFE | -30.85 |
| Energy contribution | -30.69 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740565 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
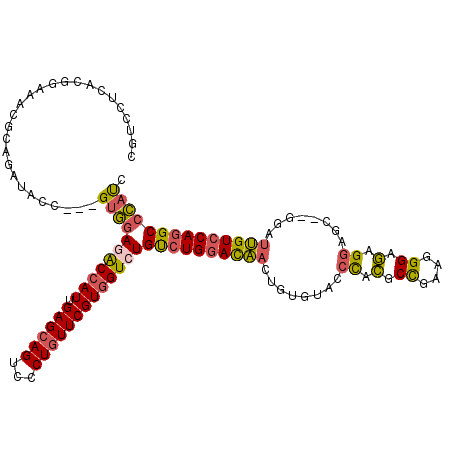
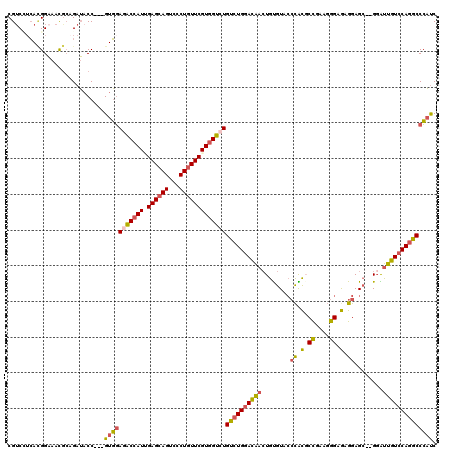
>2L_DroMel_CAF1 13133540 115 + 22407834 CGUCCUCACGGAAAUGCAGAUACC---GUGGAGACCAUAGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAACUGUGUACCCACGCCGAAGGGAGAGGAGC--GGAUUGUCCAGGCCCAUC .....((((((...........))---))))(((((((.((((((...)))))))))))))((((((((((((((....((.(.((....)).).)).))--)).))))))))))..... ( -48.30) >DroSec_CAF1 832 115 + 1 CGUCAUCACGGAAAUGCAGAUACC---GUGGAGACCAUAGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAACUGUGUACCCACGCCGAAGGGCGAGGAGC--GGAUUGUCCAGGCCCAUC .....((((((...........))---))))(((((((.((((((...)))))))))))))((((((((((((((....((.((((....)))).)).))--)).))))))))))..... ( -54.20) >DroSim_CAF1 818 115 + 1 CGUCAUCACGGAAAUGCAGAUACC---GUGGAGACCAUUGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAACUGUGUACCCACGCCGAAGGGCGAGGAGC--GGAUUGUCCAGGCCCAUC .....((((((...........))---))))(((((((.((((((...)))))))))))))((((((((((((((....((.((((....)))).)).))--)).))))))))))..... ( -54.20) >DroEre_CAF1 867 115 + 1 CGUCCGCACGGAAACGCAGAUACC---GUGGAAACCAUUGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAACUGUGUGCCCGUUCCGAAGGGAAAGGAGC--GGGUUGUCCAGGCCCAUC ..(((((.((....))..(....)---))))).(((((.((((((...)))))))))))..((((((((((((...(((((.((((....)))).)).))--)))))))))))))..... ( -50.50) >DroYak_CAF1 728 115 + 1 CGACUUCAUGGAAACGCAGAUACC---GUGGAGACCAUUGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAAGUGUGUACCCACUCUGAAGGGCGAGGAGA--GGGUUGUCCAGGCCCAUC .........(....).........---(((((((((((.((((((...)))))))))))))(((((((((.......((((.((((.........)))).--))))))))))))))))). ( -43.51) >DroPer_CAF1 26530 118 + 1 CGUCCUGAGGGCAACAACGAGUCCAUUGUCCACGCCAUCGAGCAGUCCCUCUUCGUCGUCUGCCUGGAUGACUGCGUGAAUGUGCCCA--GGAAUGGAGCUAGUGCCGUGCACGCCGCCC ..((((..(((((.((((((.....)))).(((((....(((......)))...(((((((....))))))).)))))..))))))))--)))..((.((..((((...))))...)))) ( -38.40) >consensus CGUCCUCACGGAAACGCAGAUACC___GUGGAGACCAUUGAGCAGUCCCUGUUCGUGGUCUGUCUGGACAACUGUGUACCCACGCCGAAGGGAGAGGAGC__GGAUUGUCCAGGCCCAUC ...........................(((((((((((.((((((...)))))))))))))((((((((((........((.(.((....)).).))........)))))))))))))). (-30.85 = -30.69 + -0.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 11:21:45 2006